You are viewing a plain text version of this content. The canonical link for it is here.
Posted to issues@iceberg.apache.org by GitBox <gi...@apache.org> on 2020/12/09 07:31:22 UTC
[GitHub] [iceberg] islandgit opened a new issue #1894: Spark writes the Iceberg dual partition table to report an error
islandgit opened a new issue #1894:
URL: https://github.com/apache/iceberg/issues/1894
When I write data to the table using the spark to the iceberg , table partition is time partition + bucket, the spark an error: Java. Lang. An IllegalStateException: Already closed files for partition: p_time_month = 2020-08 / vehicle_bucket = 0
Data write code:
spark.createDataFrame(sinoiov,schema)
.sortWithinPartitions("p_time","vehicle")
.write
.format("iceberg")
.mode("append")
.save("iceberg_db.original_data")
----------------------------------------------------------------
This is an automated message from the Apache Git Service.
To respond to the message, please log on to GitHub and use the
URL above to go to the specific comment.
For queries about this service, please contact Infrastructure at:
users@infra.apache.org
---------------------------------------------------------------------
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For additional commands, e-mail: issues-help@iceberg.apache.org
[GitHub] [iceberg] RussellSpitzer edited a comment on issue #1894: Spark writes the Iceberg dual partition table to report an error
Posted by GitBox <gi...@apache.org>.
RussellSpitzer edited a comment on issue #1894:
URL: https://github.com/apache/iceberg/issues/1894#issuecomment-774276183
"Already closed files" tends to occur when the write data was not actually partitioned on the correct columns. This is an error that says the data within a Spark Partition contained data for multiple Iceberg Partitions and that this data was not ordered correctly. IE there was a record for Iceberg partition A, then one for Iceberg partition B, then one for A again.
So my guess would be that `partition_by_columns` does not match the table
----------------------------------------------------------------
This is an automated message from the Apache Git Service.
To respond to the message, please log on to GitHub and use the
URL above to go to the specific comment.
For queries about this service, please contact Infrastructure at:
users@infra.apache.org
---------------------------------------------------------------------
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For additional commands, e-mail: issues-help@iceberg.apache.org
[GitHub] [iceberg] RussellSpitzer commented on issue #1894: Spark writes the Iceberg dual partition table to report an error
Posted by GitBox <gi...@apache.org>.
RussellSpitzer commented on issue #1894:
URL: https://github.com/apache/iceberg/issues/1894#issuecomment-774303754
All Iceberg tables exist in the context of a Catalog. If you haven't specified one and just provided a path it is using the implied "HadoopCatalog". The table will have It's own Metadata.json file which will specify the schema of the table and partition spec. Once created, that is the schema and partition spec until it is specifically altered.
So in your example "./data-iceberg" will be treated as a HadoopTable and look in that directory for a Metadata.json with a specified schema and partitioning. If such a file did not exist you would get an error like
```scala> spark.read.parquet("/Users/russellspitzer/Temp/parquet_data").write.format("iceberg").save("/Users/russellspitzer/Temp/iceberg_table")
org.apache.iceberg.exceptions.NoSuchTableException: Table does not exist at location: /Users/russellspitzer/Temp/iceberg_table```
Since you didn't get such an error means that a table does exist there and it has a schema.
When attempting attempting to write Spark saw "partitionBy" "a", "b", and preformed a shuffle on the input data
So all spark partitions are now like
```
SparkPartition 1 - > (1,1,1,1) , (1,1,2,1), (1,1,1,3) ...
SparkPartition 2 -> (1,2,1,1), (1,2,2,1), (1,2,1,2) ...
SparkPartition 3 -> (2,4,1,1), (2,4,2,1), (2,4,1,2) ...
```
The first two values are always the same within the spark partition, but the last two are in a random order.
Then the Iceberg writer gets one of these partitions and it creates a writer based on it's Metadata.json which let's say is not
the same as the `partitionedBy` argument. For this example let's say the table is actually partitioned by 'c'
By default in Spark Iceberg's writer opens a file for each partition assumes it will write all the values for that partition, then close the file. If it tries to open partition 1 it ends up doing the following
```
(1,1,1,1) -> Open partition c = 1 and start new file
(1,1,2,1) -> Close partition c = 1 file; Open partition c =2 and start new file
(1,1,1,3) -> Open partition c = 1 /// Error already closed file for Partition c=1
```
Since this is a runtime check you won't see this error with a single row since that will always be able to open and close the file without incident, but a dataset with multiple rows can potentially trigger this. Most of the time you want this Spark partitioning to match the Iceberg partitioning because this minimizes the number of files you need to actually write.
If you really don't want to have the Spark Partitioning match for some reason you can use the sortWithinPartitions to order the data within the partitions by their Iceberg partition values.
The other possibility is to use the new spark fanout writer which keeps all of the files open until they entire task has been completed.
Neither of these solutions is really as good as having your data partitioned like the data is expecting since that minimizes the number of files being written since all the files for a given partition will be done by the same Spark Task.
----------------------------------------------------------------
This is an automated message from the Apache Git Service.
To respond to the message, please log on to GitHub and use the
URL above to go to the specific comment.
For queries about this service, please contact Infrastructure at:
users@infra.apache.org
---------------------------------------------------------------------
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For additional commands, e-mail: issues-help@iceberg.apache.org
[GitHub] [iceberg] RussellSpitzer commented on issue #1894: Spark writes the Iceberg dual partition table to report an error
Posted by GitBox <gi...@apache.org>.
RussellSpitzer commented on issue #1894:
URL: https://github.com/apache/iceberg/issues/1894#issuecomment-774307653
And here is a example
```scala
scala> spark.sql("CREATE table ice.default.part (x Int, y Int) USING iceberg partitioned by (x) location '/Users/russellspitzer/Temp/iceberg_table'");
val df = spark.createDataFrame(Seq((1,1),(1,2),(1,3),(2,1),(2,2),(2,3),(3,1),(3,2),(3,3))).toDF("x","y")
scala> df.write.format("iceberg").partitionBy("x").mode("overwrite").saveAsTable("ice.default.part")
// No Error
scala> df.write.format("iceberg").partitionBy("y").mode("overwrite").saveAsTable("ice.default.part")
// Many errors
// Caused by: java.lang.IllegalStateException: Already closed files for partition: y=1
```
----------------------------------------------------------------
This is an automated message from the Apache Git Service.
To respond to the message, please log on to GitHub and use the
URL above to go to the specific comment.
For queries about this service, please contact Infrastructure at:
users@infra.apache.org
---------------------------------------------------------------------
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For additional commands, e-mail: issues-help@iceberg.apache.org
[GitHub] [iceberg] nchammas commented on issue #1894: Spark writes the Iceberg dual partition table to report an error
Posted by GitBox <gi...@apache.org>.
nchammas commented on issue #1894:
URL: https://github.com/apache/iceberg/issues/1894#issuecomment-774269683
I'm getting a similar exception on Spark 3.0.1 / Iceberg 0.11, though I don't understand the title of this issue, so perhaps my problem is unrelated.
I was just trying to rewrite some Parquet data as an Iceberg table:
```python
(
partition_balanced_data
.write
.format('iceberg')
.partitionBy(partition_by_columns)
.save('s3://.../exposure-iceberg/')
)
```
Here's the stack trace:
```
21/02/05 20:15:51 WARN TaskSetManager: Lost task 1151.0 in stage 14.0 (TID 295528, ip-10-0-10-131.ec2.internal, executor 1751): TaskKilled (Stage cancelled)
py4j.protocol.Py4JJavaError: An error occurred while calling o140.save.
: org.apache.spark.SparkException: Writing job aborted.
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2(WriteToDataSourceV2Exec.scala:413)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2$(WriteToDataSourceV2Exec.scala:361)
at org.apache.spark.sql.execution.datasources.v2.AtomicCreateTableAsSelectExec.writeWithV2(WriteToDataSourceV2Exec.scala:118)
at org.apache.spark.sql.execution.datasources.v2.AtomicTableWriteExec.$anonfun$writeToStagedTable$1(WriteToDataSourceV2Exec.scala:500)
at org.apache.spark.util.Utils$.tryWithSafeFinallyAndFailureCallbacks(Utils.scala:1411)
at org.apache.spark.sql.execution.datasources.v2.AtomicTableWriteExec.writeToStagedTable(WriteToDataSourceV2Exec.scala:489)
at org.apache.spark.sql.execution.datasources.v2.AtomicTableWriteExec.writeToStagedTable$(WriteToDataSourceV2Exec.scala:485)
at org.apache.spark.sql.execution.datasources.v2.AtomicCreateTableAsSelectExec.writeToStagedTable(WriteToDataSourceV2Exec.scala:118)
at org.apache.spark.sql.execution.datasources.v2.AtomicCreateTableAsSelectExec.run(WriteToDataSourceV2Exec.scala:138)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.result$lzycompute(V2CommandExec.scala:39)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.result(V2CommandExec.scala:39)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.doExecute(V2CommandExec.scala:54)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$execute$1(SparkPlan.scala:180)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$executeQuery$1(SparkPlan.scala:218)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:151)
at org.apache.spark.sql.execution.SparkPlan.executeQuery(SparkPlan.scala:215)
at org.apache.spark.sql.execution.SparkPlan.execute(SparkPlan.scala:176)
at org.apache.spark.sql.execution.QueryExecution.toRdd$lzycompute(QueryExecution.scala:124)
at org.apache.spark.sql.execution.QueryExecution.toRdd(QueryExecution.scala:123)
at org.apache.spark.sql.DataFrameWriter.$anonfun$runCommand$1(DataFrameWriter.scala:963)
at org.apache.spark.sql.catalyst.QueryPlanningTracker$.withTracker(QueryPlanningTracker.scala:104)
at org.apache.spark.sql.execution.SQLExecution$.withTracker(SQLExecution.scala:227)
at org.apache.spark.sql.execution.SQLExecution$.executeQuery$1(SQLExecution.scala:107)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$6(SQLExecution.scala:132)
at org.apache.spark.sql.catalyst.QueryPlanningTracker$.withTracker(QueryPlanningTracker.scala:104)
at org.apache.spark.sql.execution.SQLExecution$.withTracker(SQLExecution.scala:227)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$5(SQLExecution.scala:132)
at org.apache.spark.sql.execution.SQLExecution$.withSQLConfPropagated(SQLExecution.scala:248)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$1(SQLExecution.scala:131)
at org.apache.spark.sql.SparkSession.withActive(SparkSession.scala:764)
at org.apache.spark.sql.execution.SQLExecution$.withNewExecutionId(SQLExecution.scala:68)
at org.apache.spark.sql.DataFrameWriter.runCommand(DataFrameWriter.scala:963)
at org.apache.spark.sql.DataFrameWriter.save(DataFrameWriter.scala:382)
at org.apache.spark.sql.DataFrameWriter.save(DataFrameWriter.scala:288)
at sun.reflect.NativeMethodAccessorImpl.invoke0(Native Method)
at sun.reflect.NativeMethodAccessorImpl.invoke(NativeMethodAccessorImpl.java:62)
at sun.reflect.DelegatingMethodAccessorImpl.invoke(DelegatingMethodAccessorImpl.java:43)
at java.lang.reflect.Method.invoke(Method.java:498)
at py4j.reflection.MethodInvoker.invoke(MethodInvoker.java:244)
at py4j.reflection.ReflectionEngine.invoke(ReflectionEngine.java:357)
at py4j.Gateway.invoke(Gateway.java:282)
at py4j.commands.AbstractCommand.invokeMethod(AbstractCommand.java:132)
at py4j.commands.CallCommand.execute(CallCommand.java:79)
at py4j.GatewayConnection.run(GatewayConnection.java:238)
at java.lang.Thread.run(Thread.java:748)
Caused by: org.apache.spark.SparkException: Job aborted due to stage failure: Task 277 in stage 14.0 failed 4 times, most recent failure: Lost task 277.3 in stage 14.0 (TID 295505, ip-10-0-10-34.ec2.internal, executor 1935): java.lang.IllegalStateException: Already closed files for partition: exposure_data_source_id=883/exposure_date_utc=2020-12-20
at org.apache.iceberg.io.PartitionedWriter.write(PartitionedWriter.java:69)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.$anonfun$run$7(WriteToDataSourceV2Exec.scala:441)
at org.apache.spark.util.Utils$.tryWithSafeFinallyAndFailureCallbacks(Utils.scala:1411)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.run(WriteToDataSourceV2Exec.scala:477)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.$anonfun$writeWithV2$2(WriteToDataSourceV2Exec.scala:385)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:127)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:446)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1377)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:449)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
at java.lang.Thread.run(Thread.java:748)
Driver stacktrace:
at org.apache.spark.scheduler.DAGScheduler.failJobAndIndependentStages(DAGScheduler.scala:2215)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2(DAGScheduler.scala:2164)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2$adapted(DAGScheduler.scala:2163)
at scala.collection.mutable.ResizableArray.foreach(ResizableArray.scala:62)
at scala.collection.mutable.ResizableArray.foreach$(ResizableArray.scala:55)
at scala.collection.mutable.ArrayBuffer.foreach(ArrayBuffer.scala:49)
at org.apache.spark.scheduler.DAGScheduler.abortStage(DAGScheduler.scala:2163)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1(DAGScheduler.scala:1013)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1$adapted(DAGScheduler.scala:1013)
at scala.Option.foreach(Option.scala:407)
at org.apache.spark.scheduler.DAGScheduler.handleTaskSetFailed(DAGScheduler.scala:1013)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.doOnReceive(DAGScheduler.scala:2395)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2344)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2333)
at org.apache.spark.util.EventLoop$$anon$1.run(EventLoop.scala:49)
at org.apache.spark.scheduler.DAGScheduler.runJob(DAGScheduler.scala:815)
at org.apache.spark.SparkContext.runJob(SparkContext.scala:2099)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2(WriteToDataSourceV2Exec.scala:382)
... 44 more
Caused by: java.lang.IllegalStateException: Already closed files for partition: exposure_data_source_id=883/exposure_date_utc=2020-12-20
at org.apache.iceberg.io.PartitionedWriter.write(PartitionedWriter.java:69)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.$anonfun$run$7(WriteToDataSourceV2Exec.scala:441)
at org.apache.spark.util.Utils$.tryWithSafeFinallyAndFailureCallbacks(Utils.scala:1411)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.run(WriteToDataSourceV2Exec.scala:477)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.$anonfun$writeWithV2$2(WriteToDataSourceV2Exec.scala:385)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:127)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:446)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1377)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:449)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
... 1 more
```
----------------------------------------------------------------
This is an automated message from the Apache Git Service.
To respond to the message, please log on to GitHub and use the
URL above to go to the specific comment.
For queries about this service, please contact Infrastructure at:
users@infra.apache.org
---------------------------------------------------------------------
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For additional commands, e-mail: issues-help@iceberg.apache.org
[GitHub] [iceberg] Ranjith-AR edited a comment on issue #1894: Spark writes the Iceberg dual partition table to report an error
Posted by GitBox <gi...@apache.org>.
Ranjith-AR edited a comment on issue #1894:
URL: https://github.com/apache/iceberg/issues/1894#issuecomment-1050738332
I tried replicating the dual partition example provided in the docs
https://iceberg.apache.org/docs/latest/spark-writes/
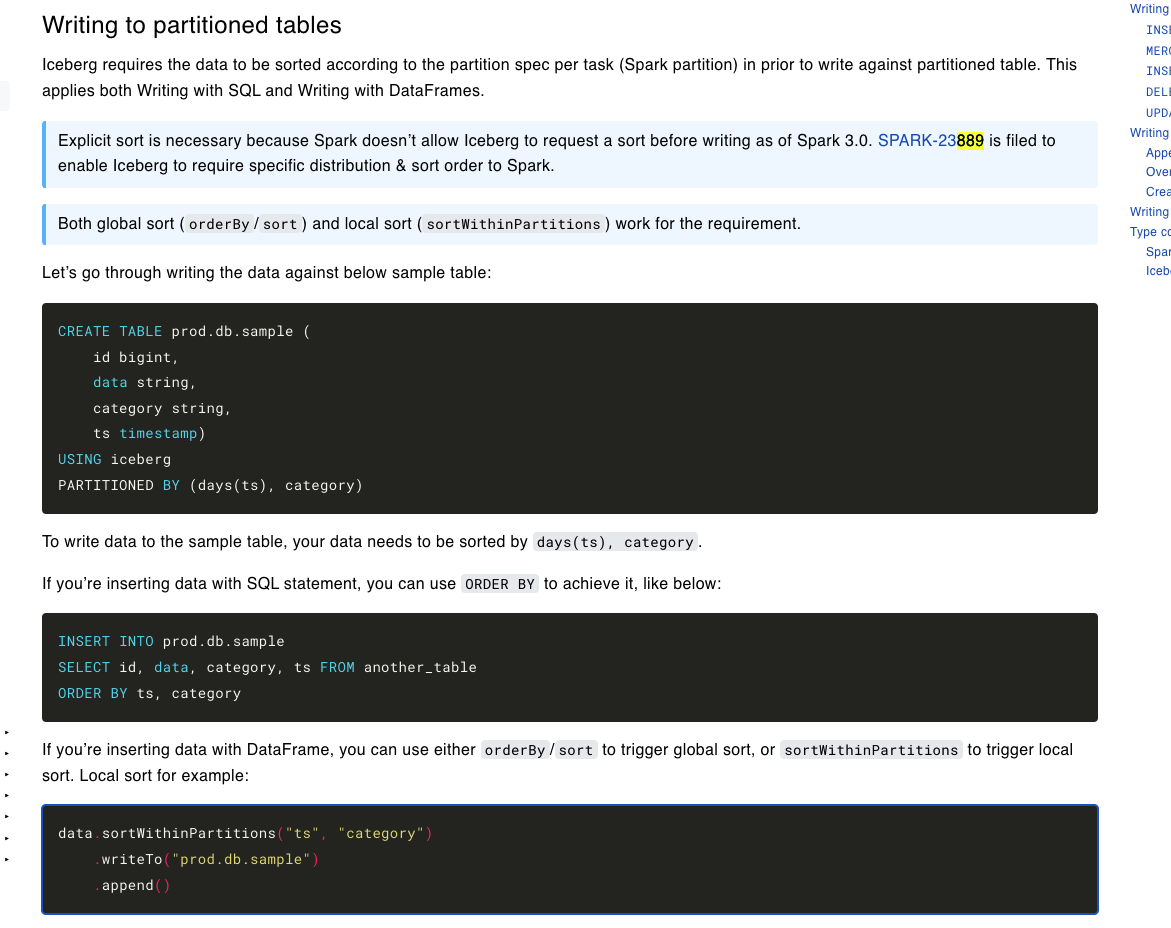
Created the below table:
-------
"CREATE TABLE nyc.green_taxis_by_days_vendorid (
VendorID string,
lpep_pickup_datetime timestamp,
lpep_dropoff_datetime string,
store_and_fwd_flag string,RatecodeID string,PULocationID string,DOLocationID string,
passenger_count string,trip_distance string,fare_amount string,extra string,mta_tax string,
tip_amount string,tolls_amount string,ehail_fee string,improvement_surcharge string,
total_amount string,
payment_type string,trip_type string,
congestion_surcharge string)
USING iceberg
PARTITIONED BY (
***days(lpep_pickup_datetime),VendorID***
)"
-----
Trying to write into the table via the dataframe api.
df_green_stage1.sortWithinPartitions(("lpep_pickup_datetime"),"VendorID") \
.writeTo("nyc.green_taxis_by_days_vendorid") \
.append()
------
However, I run into the below error
The partitions are defined as days(timestamp) and secondary as vendor id.
since the dataframe is sorted by "timestamp, vendor id" and not as "days(timestamp), vendor id" as defined in the iceberg table , we get the file already closed error.
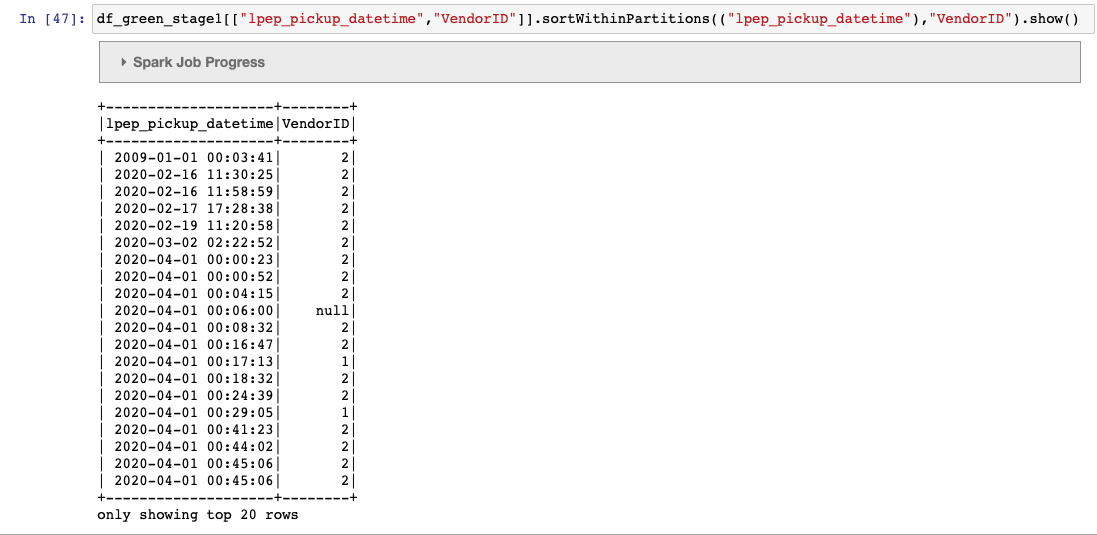
Is this a bug ? any work arounds (other than adding an additional date column and using that in the sort) ?
--------
An error was encountered:
An error occurred while calling o294.append.
: org.apache.spark.SparkException: Writing job aborted.
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2(WriteToDataSourceV2Exec.scala:388)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2$(WriteToDataSourceV2Exec.scala:336)
at org.apache.spark.sql.execution.datasources.v2.AppendDataExec.writeWithV2(WriteToDataSourceV2Exec.scala:218)
at org.apache.spark.sql.execution.datasources.v2.AppendDataExec.run(WriteToDataSourceV2Exec.scala:225)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.result$lzycompute(V2CommandExec.scala:40)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.result(V2CommandExec.scala:40)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.doExecute(V2CommandExec.scala:55)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$execute$1(SparkPlan.scala:194)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$executeQuery$1(SparkPlan.scala:232)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:151)
at org.apache.spark.sql.execution.SparkPlan.executeQuery(SparkPlan.scala:229)
at org.apache.spark.sql.execution.SparkPlan.execute(SparkPlan.scala:190)
at org.apache.spark.sql.execution.QueryExecution.toRdd$lzycompute(QueryExecution.scala:134)
at org.apache.spark.sql.execution.QueryExecution.toRdd(QueryExecution.scala:133)
at org.apache.spark.sql.DataFrameWriterV2.$anonfun$runCommand$1(DataFrameWriterV2.scala:196)
at org.apache.spark.sql.catalyst.QueryPlanningTracker$.withTracker(QueryPlanningTracker.scala:107)
at org.apache.spark.sql.execution.SQLExecution$.withTracker(SQLExecution.scala:232)
at org.apache.spark.sql.execution.SQLExecution$.executeQuery$1(SQLExecution.scala:110)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$6(SQLExecution.scala:135)
at org.apache.spark.sql.catalyst.QueryPlanningTracker$.withTracker(QueryPlanningTracker.scala:107)
at org.apache.spark.sql.execution.SQLExecution$.withTracker(SQLExecution.scala:232)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$5(SQLExecution.scala:135)
at org.apache.spark.sql.execution.SQLExecution$.withSQLConfPropagated(SQLExecution.scala:253)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$1(SQLExecution.scala:134)
at org.apache.spark.sql.SparkSession.withActive(SparkSession.scala:775)
at org.apache.spark.sql.execution.SQLExecution$.withNewExecutionId(SQLExecution.scala:68)
at org.apache.spark.sql.DataFrameWriterV2.runCommand(DataFrameWriterV2.scala:196)
at org.apache.spark.sql.DataFrameWriterV2.append(DataFrameWriterV2.scala:149)
at sun.reflect.NativeMethodAccessorImpl.invoke0(Native Method)
at sun.reflect.NativeMethodAccessorImpl.invoke(NativeMethodAccessorImpl.java:62)
at sun.reflect.DelegatingMethodAccessorImpl.invoke(DelegatingMethodAccessorImpl.java:43)
at java.lang.reflect.Method.invoke(Method.java:498)
at py4j.reflection.MethodInvoker.invoke(MethodInvoker.java:244)
at py4j.reflection.ReflectionEngine.invoke(ReflectionEngine.java:357)
at py4j.Gateway.invoke(Gateway.java:282)
at py4j.commands.AbstractCommand.invokeMethod(AbstractCommand.java:132)
at py4j.commands.CallCommand.execute(CallCommand.java:79)
at py4j.GatewayConnection.run(GatewayConnection.java:238)
at java.lang.Thread.run(Thread.java:750)
Caused by: org.apache.spark.SparkException: Job aborted due to stage failure: Task 0 in stage 32.0 failed 4 times, most recent failure: Lost task 0.3 in stage 32.0 (TID 103) (ip-172-31-21-243.ec2.internal executor 18): java.lang.IllegalStateException: Incoming records violate the writer assumption that records are clustered by spec and by partition within each spec. Either cluster the incoming records or switch to fanout writers.
Encountered records that belong to already closed files:
partition 'lpep_pickup_datetime_day=2020-04-01/VendorID=2' in spec [
1000: lpep_pickup_datetime_day: day(2)
1001: VendorID: identity(1)
]
at org.apache.iceberg.io.ClusteredWriter.write(ClusteredWriter.java:95)
at org.apache.iceberg.io.ClusteredDataWriter.write(ClusteredDataWriter.java:34)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:629)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:604)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.$anonfun$run$1(WriteToDataSourceV2Exec.scala:416)
at org.apache.spark.util.Utils$.tryWithSafeFinallyAndFailureCallbacks(Utils.scala:1473)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.run(WriteToDataSourceV2Exec.scala:452)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.$anonfun$writeWithV2$2(WriteToDataSourceV2Exec.scala:360)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:131)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:497)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1439)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:500)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
at java.lang.Thread.run(Thread.java:750)
Driver stacktrace:
at org.apache.spark.scheduler.DAGScheduler.failJobAndIndependentStages(DAGScheduler.scala:2470)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2(DAGScheduler.scala:2419)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2$adapted(DAGScheduler.scala:2418)
at scala.collection.mutable.ResizableArray.foreach(ResizableArray.scala:62)
at scala.collection.mutable.ResizableArray.foreach$(ResizableArray.scala:55)
at scala.collection.mutable.ArrayBuffer.foreach(ArrayBuffer.scala:49)
at org.apache.spark.scheduler.DAGScheduler.abortStage(DAGScheduler.scala:2418)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1(DAGScheduler.scala:1125)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1$adapted(DAGScheduler.scala:1125)
at scala.Option.foreach(Option.scala:407)
at org.apache.spark.scheduler.DAGScheduler.handleTaskSetFailed(DAGScheduler.scala:1125)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.doOnReceive(DAGScheduler.scala:2684)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2626)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2615)
at org.apache.spark.util.EventLoop$$anon$1.run(EventLoop.scala:49)
at org.apache.spark.scheduler.DAGScheduler.runJob(DAGScheduler.scala:914)
at org.apache.spark.SparkContext.runJob(SparkContext.scala:2241)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2(WriteToDataSourceV2Exec.scala:357)
... 38 more
Caused by: java.lang.IllegalStateException: Incoming records violate the writer assumption that records are clustered by spec and by partition within each spec. Either cluster the incoming records or switch to fanout writers.
Encountered records that belong to already closed files:
partition 'lpep_pickup_datetime_day=2020-04-01/VendorID=2' in spec [
1000: lpep_pickup_datetime_day: day(2)
1001: VendorID: identity(1)
]
at org.apache.iceberg.io.ClusteredWriter.write(ClusteredWriter.java:95)
at org.apache.iceberg.io.ClusteredDataWriter.write(ClusteredDataWriter.java:34)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:629)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:604)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.$anonfun$run$1(WriteToDataSourceV2Exec.scala:416)
at org.apache.spark.util.Utils$.tryWithSafeFinallyAndFailureCallbacks(Utils.scala:1473)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.run(WriteToDataSourceV2Exec.scala:452)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.$anonfun$writeWithV2$2(WriteToDataSourceV2Exec.scala:360)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:131)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:497)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1439)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:500)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
... 1 more
Traceback (most recent call last):
File "/usr/lib/spark/python/lib/pyspark.zip/pyspark/sql/readwriter.py", line 1564, in append
self._jwriter.append()
File "/usr/lib/spark/python/lib/py4j-0.10.9-src.zip/py4j/java_gateway.py", line 1305, in __call__
answer, self.gateway_client, self.target_id, self.name)
File "/usr/lib/spark/python/lib/pyspark.zip/pyspark/sql/utils.py", line 111, in deco
return f(*a, **kw)
File "/usr/lib/spark/python/lib/py4j-0.10.9-src.zip/py4j/protocol.py", line 328, in get_return_value
format(target_id, ".", name), value)
py4j.protocol.Py4JJavaError: An error occurred while calling o294.append.
: org.apache.spark.SparkException: Writing job aborted.
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2(WriteToDataSourceV2Exec.scala:388)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2$(WriteToDataSourceV2Exec.scala:336)
at org.apache.spark.sql.execution.datasources.v2.AppendDataExec.writeWithV2(WriteToDataSourceV2Exec.scala:218)
at org.apache.spark.sql.execution.datasources.v2.AppendDataExec.run(WriteToDataSourceV2Exec.scala:225)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.result$lzycompute(V2CommandExec.scala:40)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.result(V2CommandExec.scala:40)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.doExecute(V2CommandExec.scala:55)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$execute$1(SparkPlan.scala:194)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$executeQuery$1(SparkPlan.scala:232)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:151)
at org.apache.spark.sql.execution.SparkPlan.executeQuery(SparkPlan.scala:229)
at org.apache.spark.sql.execution.SparkPlan.execute(SparkPlan.scala:190)
at org.apache.spark.sql.execution.QueryExecution.toRdd$lzycompute(QueryExecution.scala:134)
at org.apache.spark.sql.execution.QueryExecution.toRdd(QueryExecution.scala:133)
at org.apache.spark.sql.DataFrameWriterV2.$anonfun$runCommand$1(DataFrameWriterV2.scala:196)
at org.apache.spark.sql.catalyst.QueryPlanningTracker$.withTracker(QueryPlanningTracker.scala:107)
at org.apache.spark.sql.execution.SQLExecution$.withTracker(SQLExecution.scala:232)
at org.apache.spark.sql.execution.SQLExecution$.executeQuery$1(SQLExecution.scala:110)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$6(SQLExecution.scala:135)
at org.apache.spark.sql.catalyst.QueryPlanningTracker$.withTracker(QueryPlanningTracker.scala:107)
at org.apache.spark.sql.execution.SQLExecution$.withTracker(SQLExecution.scala:232)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$5(SQLExecution.scala:135)
at org.apache.spark.sql.execution.SQLExecution$.withSQLConfPropagated(SQLExecution.scala:253)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$1(SQLExecution.scala:134)
at org.apache.spark.sql.SparkSession.withActive(SparkSession.scala:775)
at org.apache.spark.sql.execution.SQLExecution$.withNewExecutionId(SQLExecution.scala:68)
at org.apache.spark.sql.DataFrameWriterV2.runCommand(DataFrameWriterV2.scala:196)
at org.apache.spark.sql.DataFrameWriterV2.append(DataFrameWriterV2.scala:149)
at sun.reflect.NativeMethodAccessorImpl.invoke0(Native Method)
at sun.reflect.NativeMethodAccessorImpl.invoke(NativeMethodAccessorImpl.java:62)
at sun.reflect.DelegatingMethodAccessorImpl.invoke(DelegatingMethodAccessorImpl.java:43)
at java.lang.reflect.Method.invoke(Method.java:498)
at py4j.reflection.MethodInvoker.invoke(MethodInvoker.java:244)
at py4j.reflection.ReflectionEngine.invoke(ReflectionEngine.java:357)
at py4j.Gateway.invoke(Gateway.java:282)
at py4j.commands.AbstractCommand.invokeMethod(AbstractCommand.java:132)
at py4j.commands.CallCommand.execute(CallCommand.java:79)
at py4j.GatewayConnection.run(GatewayConnection.java:238)
at java.lang.Thread.run(Thread.java:750)
Caused by: org.apache.spark.SparkException: Job aborted due to stage failure: Task 0 in stage 32.0 failed 4 times, most recent failure: Lost task 0.3 in stage 32.0 (TID 103) (ip-172-31-21-243.ec2.internal executor 18): java.lang.IllegalStateException: Incoming records violate the writer assumption that records are clustered by spec and by partition within each spec. Either cluster the incoming records or switch to fanout writers.
Encountered records that belong to already closed files:
partition 'lpep_pickup_datetime_day=2020-04-01/VendorID=2' in spec [
1000: lpep_pickup_datetime_day: day(2)
1001: VendorID: identity(1)
]
at org.apache.iceberg.io.ClusteredWriter.write(ClusteredWriter.java:95)
at org.apache.iceberg.io.ClusteredDataWriter.write(ClusteredDataWriter.java:34)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:629)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:604)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.$anonfun$run$1(WriteToDataSourceV2Exec.scala:416)
at org.apache.spark.util.Utils$.tryWithSafeFinallyAndFailureCallbacks(Utils.scala:1473)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.run(WriteToDataSourceV2Exec.scala:452)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.$anonfun$writeWithV2$2(WriteToDataSourceV2Exec.scala:360)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:131)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:497)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1439)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:500)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
at java.lang.Thread.run(Thread.java:750)
Driver stacktrace:
at org.apache.spark.scheduler.DAGScheduler.failJobAndIndependentStages(DAGScheduler.scala:2470)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2(DAGScheduler.scala:2419)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2$adapted(DAGScheduler.scala:2418)
at scala.collection.mutable.ResizableArray.foreach(ResizableArray.scala:62)
at scala.collection.mutable.ResizableArray.foreach$(ResizableArray.scala:55)
at scala.collection.mutable.ArrayBuffer.foreach(ArrayBuffer.scala:49)
at org.apache.spark.scheduler.DAGScheduler.abortStage(DAGScheduler.scala:2418)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1(DAGScheduler.scala:1125)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1$adapted(DAGScheduler.scala:1125)
at scala.Option.foreach(Option.scala:407)
at org.apache.spark.scheduler.DAGScheduler.handleTaskSetFailed(DAGScheduler.scala:1125)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.doOnReceive(DAGScheduler.scala:2684)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2626)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2615)
at org.apache.spark.util.EventLoop$$anon$1.run(EventLoop.scala:49)
at org.apache.spark.scheduler.DAGScheduler.runJob(DAGScheduler.scala:914)
at org.apache.spark.SparkContext.runJob(SparkContext.scala:2241)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2(WriteToDataSourceV2Exec.scala:357)
... 38 more
Caused by: java.lang.IllegalStateException: Incoming records violate the writer assumption that records are clustered by spec and by partition within each spec. Either cluster the incoming records or switch to fanout writers.
Encountered records that belong to already closed files:
partition 'lpep_pickup_datetime_day=2020-04-01/VendorID=2' in spec [
1000: lpep_pickup_datetime_day: day(2)
1001: VendorID: identity(1)
]
at org.apache.iceberg.io.ClusteredWriter.write(ClusteredWriter.java:95)
at org.apache.iceberg.io.ClusteredDataWriter.write(ClusteredDataWriter.java:34)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:629)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:604)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.$anonfun$run$1(WriteToDataSourceV2Exec.scala:416)
at org.apache.spark.util.Utils$.tryWithSafeFinallyAndFailureCallbacks(Utils.scala:1473)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.run(WriteToDataSourceV2Exec.scala:452)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.$anonfun$writeWithV2$2(WriteToDataSourceV2Exec.scala:360)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:131)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:497)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1439)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:500)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
... 1 more
-------
--
This is an automated message from the Apache Git Service.
To respond to the message, please log on to GitHub and use the
URL above to go to the specific comment.
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For queries about this service, please contact Infrastructure at:
users@infra.apache.org
---------------------------------------------------------------------
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For additional commands, e-mail: issues-help@iceberg.apache.org
[GitHub] [iceberg] pan3793 commented on issue #1894: Spark writes the Iceberg dual partition table to report an error
Posted by GitBox <gi...@apache.org>.
pan3793 commented on issue #1894:
URL: https://github.com/apache/iceberg/issues/1894#issuecomment-1014572605
I think #3461 should address this issue, but it's only for Spark 3.2, does the community have a plan(or at least accept backport PR) for Spark 3.1? cc @RussellSpitzer @aokolnychyi
--
This is an automated message from the Apache Git Service.
To respond to the message, please log on to GitHub and use the
URL above to go to the specific comment.
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For queries about this service, please contact Infrastructure at:
users@infra.apache.org
---------------------------------------------------------------------
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For additional commands, e-mail: issues-help@iceberg.apache.org
[GitHub] [iceberg] Ranjith-AR edited a comment on issue #1894: Spark writes the Iceberg dual partition table to report an error
Posted by GitBox <gi...@apache.org>.
Ranjith-AR edited a comment on issue #1894:
URL: https://github.com/apache/iceberg/issues/1894#issuecomment-1050738332
I tried replicating the dual partition example provided in the docs
https://iceberg.apache.org/docs/latest/spark-writes/

Created the below table:
-------
"CREATE TABLE nyc.green_taxis_by_days_vendorid (
VendorID string,
lpep_pickup_datetime timestamp,
lpep_dropoff_datetime string,
store_and_fwd_flag string,RatecodeID string,PULocationID string,DOLocationID string,
passenger_count string,trip_distance string,fare_amount string,extra string,mta_tax string,
tip_amount string,tolls_amount string,ehail_fee string,improvement_surcharge string,
total_amount string,
payment_type string,trip_type string,
congestion_surcharge string)
USING iceberg
PARTITIONED BY (
***days(lpep_pickup_datetime),VendorID***
)"
-----
Trying to write into the table via both dataframe api and pyspark sql...and get the same error (i.e. file already closed)
df_green_stage1.sortWithinPartitions(("lpep_pickup_datetime"),"VendorID") \
.writeTo("nyc.green_taxis_by_days_vendorid") \
.append()
-----
%%sql
INSERT INTO nyc.green_taxis_by_days_vendorid (
SELECT
VendorID,lpep_pickup_datetime,lpep_dropoff_datetime,store_and_fwd_flag,RatecodeID,PULocationID,DOLocationID,passenger_count,trip_distance,fare_amount,
extra,mta_tax,tip_amount,tolls_amount,ehail_fee,
improvement_surcharge,total_amount,payment_type,trip_type,congestion_surcharge
FROM nyc.green_taxis_by_days_vendorid_withdate
ORDER BY lpep_pickup_datetime,VendorID
)
------
However, I run into the below error
The partitions are defined as days(timestamp) and secondary as vendor id.
since the dataframe is sorted by "timestamp, vendor id" and not as "days(timestamp), vendor id" as defined in the iceberg table , we get the file already closed error.

Is this a bug ? any work arounds (other than adding an additional date column and using that in the sort) ?
--------
An error was encountered:
An error occurred while calling o294.append.
: org.apache.spark.SparkException: Writing job aborted.
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2(WriteToDataSourceV2Exec.scala:388)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2$(WriteToDataSourceV2Exec.scala:336)
at org.apache.spark.sql.execution.datasources.v2.AppendDataExec.writeWithV2(WriteToDataSourceV2Exec.scala:218)
at org.apache.spark.sql.execution.datasources.v2.AppendDataExec.run(WriteToDataSourceV2Exec.scala:225)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.result$lzycompute(V2CommandExec.scala:40)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.result(V2CommandExec.scala:40)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.doExecute(V2CommandExec.scala:55)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$execute$1(SparkPlan.scala:194)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$executeQuery$1(SparkPlan.scala:232)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:151)
at org.apache.spark.sql.execution.SparkPlan.executeQuery(SparkPlan.scala:229)
at org.apache.spark.sql.execution.SparkPlan.execute(SparkPlan.scala:190)
at org.apache.spark.sql.execution.QueryExecution.toRdd$lzycompute(QueryExecution.scala:134)
at org.apache.spark.sql.execution.QueryExecution.toRdd(QueryExecution.scala:133)
at org.apache.spark.sql.DataFrameWriterV2.$anonfun$runCommand$1(DataFrameWriterV2.scala:196)
at org.apache.spark.sql.catalyst.QueryPlanningTracker$.withTracker(QueryPlanningTracker.scala:107)
at org.apache.spark.sql.execution.SQLExecution$.withTracker(SQLExecution.scala:232)
at org.apache.spark.sql.execution.SQLExecution$.executeQuery$1(SQLExecution.scala:110)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$6(SQLExecution.scala:135)
at org.apache.spark.sql.catalyst.QueryPlanningTracker$.withTracker(QueryPlanningTracker.scala:107)
at org.apache.spark.sql.execution.SQLExecution$.withTracker(SQLExecution.scala:232)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$5(SQLExecution.scala:135)
at org.apache.spark.sql.execution.SQLExecution$.withSQLConfPropagated(SQLExecution.scala:253)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$1(SQLExecution.scala:134)
at org.apache.spark.sql.SparkSession.withActive(SparkSession.scala:775)
at org.apache.spark.sql.execution.SQLExecution$.withNewExecutionId(SQLExecution.scala:68)
at org.apache.spark.sql.DataFrameWriterV2.runCommand(DataFrameWriterV2.scala:196)
at org.apache.spark.sql.DataFrameWriterV2.append(DataFrameWriterV2.scala:149)
at sun.reflect.NativeMethodAccessorImpl.invoke0(Native Method)
at sun.reflect.NativeMethodAccessorImpl.invoke(NativeMethodAccessorImpl.java:62)
at sun.reflect.DelegatingMethodAccessorImpl.invoke(DelegatingMethodAccessorImpl.java:43)
at java.lang.reflect.Method.invoke(Method.java:498)
at py4j.reflection.MethodInvoker.invoke(MethodInvoker.java:244)
at py4j.reflection.ReflectionEngine.invoke(ReflectionEngine.java:357)
at py4j.Gateway.invoke(Gateway.java:282)
at py4j.commands.AbstractCommand.invokeMethod(AbstractCommand.java:132)
at py4j.commands.CallCommand.execute(CallCommand.java:79)
at py4j.GatewayConnection.run(GatewayConnection.java:238)
at java.lang.Thread.run(Thread.java:750)
Caused by: org.apache.spark.SparkException: Job aborted due to stage failure: Task 0 in stage 32.0 failed 4 times, most recent failure: Lost task 0.3 in stage 32.0 (TID 103) (ip-172-31-21-243.ec2.internal executor 18): java.lang.IllegalStateException: Incoming records violate the writer assumption that records are clustered by spec and by partition within each spec. Either cluster the incoming records or switch to fanout writers.
Encountered records that belong to already closed files:
partition 'lpep_pickup_datetime_day=2020-04-01/VendorID=2' in spec [
1000: lpep_pickup_datetime_day: day(2)
1001: VendorID: identity(1)
]
at org.apache.iceberg.io.ClusteredWriter.write(ClusteredWriter.java:95)
at org.apache.iceberg.io.ClusteredDataWriter.write(ClusteredDataWriter.java:34)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:629)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:604)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.$anonfun$run$1(WriteToDataSourceV2Exec.scala:416)
at org.apache.spark.util.Utils$.tryWithSafeFinallyAndFailureCallbacks(Utils.scala:1473)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.run(WriteToDataSourceV2Exec.scala:452)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.$anonfun$writeWithV2$2(WriteToDataSourceV2Exec.scala:360)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:131)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:497)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1439)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:500)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
at java.lang.Thread.run(Thread.java:750)
Driver stacktrace:
at org.apache.spark.scheduler.DAGScheduler.failJobAndIndependentStages(DAGScheduler.scala:2470)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2(DAGScheduler.scala:2419)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2$adapted(DAGScheduler.scala:2418)
at scala.collection.mutable.ResizableArray.foreach(ResizableArray.scala:62)
at scala.collection.mutable.ResizableArray.foreach$(ResizableArray.scala:55)
at scala.collection.mutable.ArrayBuffer.foreach(ArrayBuffer.scala:49)
at org.apache.spark.scheduler.DAGScheduler.abortStage(DAGScheduler.scala:2418)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1(DAGScheduler.scala:1125)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1$adapted(DAGScheduler.scala:1125)
at scala.Option.foreach(Option.scala:407)
at org.apache.spark.scheduler.DAGScheduler.handleTaskSetFailed(DAGScheduler.scala:1125)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.doOnReceive(DAGScheduler.scala:2684)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2626)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2615)
at org.apache.spark.util.EventLoop$$anon$1.run(EventLoop.scala:49)
at org.apache.spark.scheduler.DAGScheduler.runJob(DAGScheduler.scala:914)
at org.apache.spark.SparkContext.runJob(SparkContext.scala:2241)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2(WriteToDataSourceV2Exec.scala:357)
... 38 more
Caused by: java.lang.IllegalStateException: Incoming records violate the writer assumption that records are clustered by spec and by partition within each spec. Either cluster the incoming records or switch to fanout writers.
Encountered records that belong to already closed files:
partition 'lpep_pickup_datetime_day=2020-04-01/VendorID=2' in spec [
1000: lpep_pickup_datetime_day: day(2)
1001: VendorID: identity(1)
]
at org.apache.iceberg.io.ClusteredWriter.write(ClusteredWriter.java:95)
at org.apache.iceberg.io.ClusteredDataWriter.write(ClusteredDataWriter.java:34)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:629)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:604)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.$anonfun$run$1(WriteToDataSourceV2Exec.scala:416)
at org.apache.spark.util.Utils$.tryWithSafeFinallyAndFailureCallbacks(Utils.scala:1473)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.run(WriteToDataSourceV2Exec.scala:452)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.$anonfun$writeWithV2$2(WriteToDataSourceV2Exec.scala:360)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:131)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:497)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1439)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:500)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
... 1 more
Traceback (most recent call last):
File "/usr/lib/spark/python/lib/pyspark.zip/pyspark/sql/readwriter.py", line 1564, in append
self._jwriter.append()
File "/usr/lib/spark/python/lib/py4j-0.10.9-src.zip/py4j/java_gateway.py", line 1305, in __call__
answer, self.gateway_client, self.target_id, self.name)
File "/usr/lib/spark/python/lib/pyspark.zip/pyspark/sql/utils.py", line 111, in deco
return f(*a, **kw)
File "/usr/lib/spark/python/lib/py4j-0.10.9-src.zip/py4j/protocol.py", line 328, in get_return_value
format(target_id, ".", name), value)
py4j.protocol.Py4JJavaError: An error occurred while calling o294.append.
: org.apache.spark.SparkException: Writing job aborted.
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2(WriteToDataSourceV2Exec.scala:388)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2$(WriteToDataSourceV2Exec.scala:336)
at org.apache.spark.sql.execution.datasources.v2.AppendDataExec.writeWithV2(WriteToDataSourceV2Exec.scala:218)
at org.apache.spark.sql.execution.datasources.v2.AppendDataExec.run(WriteToDataSourceV2Exec.scala:225)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.result$lzycompute(V2CommandExec.scala:40)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.result(V2CommandExec.scala:40)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.doExecute(V2CommandExec.scala:55)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$execute$1(SparkPlan.scala:194)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$executeQuery$1(SparkPlan.scala:232)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:151)
at org.apache.spark.sql.execution.SparkPlan.executeQuery(SparkPlan.scala:229)
at org.apache.spark.sql.execution.SparkPlan.execute(SparkPlan.scala:190)
at org.apache.spark.sql.execution.QueryExecution.toRdd$lzycompute(QueryExecution.scala:134)
at org.apache.spark.sql.execution.QueryExecution.toRdd(QueryExecution.scala:133)
at org.apache.spark.sql.DataFrameWriterV2.$anonfun$runCommand$1(DataFrameWriterV2.scala:196)
at org.apache.spark.sql.catalyst.QueryPlanningTracker$.withTracker(QueryPlanningTracker.scala:107)
at org.apache.spark.sql.execution.SQLExecution$.withTracker(SQLExecution.scala:232)
at org.apache.spark.sql.execution.SQLExecution$.executeQuery$1(SQLExecution.scala:110)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$6(SQLExecution.scala:135)
at org.apache.spark.sql.catalyst.QueryPlanningTracker$.withTracker(QueryPlanningTracker.scala:107)
at org.apache.spark.sql.execution.SQLExecution$.withTracker(SQLExecution.scala:232)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$5(SQLExecution.scala:135)
at org.apache.spark.sql.execution.SQLExecution$.withSQLConfPropagated(SQLExecution.scala:253)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$1(SQLExecution.scala:134)
at org.apache.spark.sql.SparkSession.withActive(SparkSession.scala:775)
at org.apache.spark.sql.execution.SQLExecution$.withNewExecutionId(SQLExecution.scala:68)
at org.apache.spark.sql.DataFrameWriterV2.runCommand(DataFrameWriterV2.scala:196)
at org.apache.spark.sql.DataFrameWriterV2.append(DataFrameWriterV2.scala:149)
at sun.reflect.NativeMethodAccessorImpl.invoke0(Native Method)
at sun.reflect.NativeMethodAccessorImpl.invoke(NativeMethodAccessorImpl.java:62)
at sun.reflect.DelegatingMethodAccessorImpl.invoke(DelegatingMethodAccessorImpl.java:43)
at java.lang.reflect.Method.invoke(Method.java:498)
at py4j.reflection.MethodInvoker.invoke(MethodInvoker.java:244)
at py4j.reflection.ReflectionEngine.invoke(ReflectionEngine.java:357)
at py4j.Gateway.invoke(Gateway.java:282)
at py4j.commands.AbstractCommand.invokeMethod(AbstractCommand.java:132)
at py4j.commands.CallCommand.execute(CallCommand.java:79)
at py4j.GatewayConnection.run(GatewayConnection.java:238)
at java.lang.Thread.run(Thread.java:750)
Caused by: org.apache.spark.SparkException: Job aborted due to stage failure: Task 0 in stage 32.0 failed 4 times, most recent failure: Lost task 0.3 in stage 32.0 (TID 103) (ip-172-31-21-243.ec2.internal executor 18): java.lang.IllegalStateException: Incoming records violate the writer assumption that records are clustered by spec and by partition within each spec. Either cluster the incoming records or switch to fanout writers.
Encountered records that belong to already closed files:
partition 'lpep_pickup_datetime_day=2020-04-01/VendorID=2' in spec [
1000: lpep_pickup_datetime_day: day(2)
1001: VendorID: identity(1)
]
at org.apache.iceberg.io.ClusteredWriter.write(ClusteredWriter.java:95)
at org.apache.iceberg.io.ClusteredDataWriter.write(ClusteredDataWriter.java:34)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:629)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:604)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.$anonfun$run$1(WriteToDataSourceV2Exec.scala:416)
at org.apache.spark.util.Utils$.tryWithSafeFinallyAndFailureCallbacks(Utils.scala:1473)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.run(WriteToDataSourceV2Exec.scala:452)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.$anonfun$writeWithV2$2(WriteToDataSourceV2Exec.scala:360)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:131)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:497)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1439)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:500)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
at java.lang.Thread.run(Thread.java:750)
Driver stacktrace:
at org.apache.spark.scheduler.DAGScheduler.failJobAndIndependentStages(DAGScheduler.scala:2470)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2(DAGScheduler.scala:2419)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2$adapted(DAGScheduler.scala:2418)
at scala.collection.mutable.ResizableArray.foreach(ResizableArray.scala:62)
at scala.collection.mutable.ResizableArray.foreach$(ResizableArray.scala:55)
at scala.collection.mutable.ArrayBuffer.foreach(ArrayBuffer.scala:49)
at org.apache.spark.scheduler.DAGScheduler.abortStage(DAGScheduler.scala:2418)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1(DAGScheduler.scala:1125)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1$adapted(DAGScheduler.scala:1125)
at scala.Option.foreach(Option.scala:407)
at org.apache.spark.scheduler.DAGScheduler.handleTaskSetFailed(DAGScheduler.scala:1125)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.doOnReceive(DAGScheduler.scala:2684)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2626)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2615)
at org.apache.spark.util.EventLoop$$anon$1.run(EventLoop.scala:49)
at org.apache.spark.scheduler.DAGScheduler.runJob(DAGScheduler.scala:914)
at org.apache.spark.SparkContext.runJob(SparkContext.scala:2241)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2(WriteToDataSourceV2Exec.scala:357)
... 38 more
Caused by: java.lang.IllegalStateException: Incoming records violate the writer assumption that records are clustered by spec and by partition within each spec. Either cluster the incoming records or switch to fanout writers.
Encountered records that belong to already closed files:
partition 'lpep_pickup_datetime_day=2020-04-01/VendorID=2' in spec [
1000: lpep_pickup_datetime_day: day(2)
1001: VendorID: identity(1)
]
at org.apache.iceberg.io.ClusteredWriter.write(ClusteredWriter.java:95)
at org.apache.iceberg.io.ClusteredDataWriter.write(ClusteredDataWriter.java:34)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:629)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:604)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.$anonfun$run$1(WriteToDataSourceV2Exec.scala:416)
at org.apache.spark.util.Utils$.tryWithSafeFinallyAndFailureCallbacks(Utils.scala:1473)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.run(WriteToDataSourceV2Exec.scala:452)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.$anonfun$writeWithV2$2(WriteToDataSourceV2Exec.scala:360)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:131)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:497)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1439)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:500)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
... 1 more
-------
--
This is an automated message from the Apache Git Service.
To respond to the message, please log on to GitHub and use the
URL above to go to the specific comment.
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For queries about this service, please contact Infrastructure at:
users@infra.apache.org
---------------------------------------------------------------------
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For additional commands, e-mail: issues-help@iceberg.apache.org
[GitHub] [iceberg] nchammas commented on issue #1894: Spark writes the Iceberg dual partition table to report an error
Posted by GitBox <gi...@apache.org>.
nchammas commented on issue #1894:
URL: https://github.com/apache/iceberg/issues/1894#issuecomment-774313769
Thank you for the detailed examples. I think I understand now. I guess this issue becomes transparent to the user once [SPARK-23889](https://issues.apache.org/jira/browse/SPARK-23889) is resolved, as called out in the docs you linked to.
There is one thing you said I am still confused about though:
> Since you didn't get such an error means that a table does exist there and it has a schema.
For some reason, I am able to save to previously non-existent Iceberg locations just fine:
```
$ pyspark --packages org.apache.iceberg:iceberg-spark3-runtime:0.11.0
Welcome to
____ __
/ __/__ ___ _____/ /__
_\ \/ _ \/ _ `/ __/ '_/
/__ / .__/\_,_/_/ /_/\_\ version 3.0.1
/_/
Using Python version 3.9.1 (default, Feb 3 2021 07:38:02)
>>> import uuid
>>> spark.range(3).write.format('iceberg').save('./' + str(uuid.uuid4()))
$ ll
drwxr-xr-x - user 5 Feb 16:59 0f7e86eb-c8d8-467b-9499-845fa90acbe7
$ ll 0f7e86eb-c8d8-467b-9499-845fa90acbe7/
drwxr-xr-x - user 5 Feb 16:59 data
drwxr-xr-x - user 5 Feb 16:59 metadata
```
Separate issue, but I just wanted to point that out.
----------------------------------------------------------------
This is an automated message from the Apache Git Service.
To respond to the message, please log on to GitHub and use the
URL above to go to the specific comment.
For queries about this service, please contact Infrastructure at:
users@infra.apache.org
---------------------------------------------------------------------
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For additional commands, e-mail: issues-help@iceberg.apache.org
[GitHub] [iceberg] RussellSpitzer commented on issue #1894: Spark writes the Iceberg dual partition table to report an error
Posted by GitBox <gi...@apache.org>.
RussellSpitzer commented on issue #1894:
URL: https://github.com/apache/iceberg/issues/1894#issuecomment-774276183
Already Close File tends to be that the data was not actually partitioned on the correct columns
----------------------------------------------------------------
This is an automated message from the Apache Git Service.
To respond to the message, please log on to GitHub and use the
URL above to go to the specific comment.
For queries about this service, please contact Infrastructure at:
users@infra.apache.org
---------------------------------------------------------------------
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For additional commands, e-mail: issues-help@iceberg.apache.org
[GitHub] [iceberg] nchammas commented on issue #1894: Spark writes the Iceberg dual partition table to report an error
Posted by GitBox <gi...@apache.org>.
nchammas commented on issue #1894:
URL: https://github.com/apache/iceberg/issues/1894#issuecomment-774284421
Hmm, I guess my mental model of how Iceberg works is incorrect.
What does it mean for the `partitionBy()` columns to "not match the table"? I thought that `partitionBy()` was an output directive, allowing me to read in a table partitioned one way and write it out partitioned a different way.
I'm not using any catalogs here. The `partition_balanced_data` DataFrame was read directly from an S3 path. There are 3 partitioning columns on the original data. I am attempting to write this data out to a separate location on S3 with only 2 partitioning columns.
So the partitioning columns on the output _do_ differ from the source table, but I'm confused as to why this would matter to Iceberg.
In other words, what's wrong with this contrived example?
```python
data = spark.createDataFrame([(1, 2, 3, 4)], schema=['a', 'b', 'c', 'd'])
data.write.partitionBy(['a', 'b', 'c']).parquet('./data')
data = spark.read.parquet('./data')
data.write.format('iceberg').partitionBy(['a', 'b']).save('./data-iceberg')
```
This example code actually works, but I am understanding from your comment that there may be something incorrect about it. This is roughly what I was trying to do when I got the stack trace I posted above.
----------------------------------------------------------------
This is an automated message from the Apache Git Service.
To respond to the message, please log on to GitHub and use the
URL above to go to the specific comment.
For queries about this service, please contact Infrastructure at:
users@infra.apache.org
---------------------------------------------------------------------
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For additional commands, e-mail: issues-help@iceberg.apache.org
[GitHub] [iceberg] RussellSpitzer edited a comment on issue #1894: Spark writes the Iceberg dual partition table to report an error
Posted by GitBox <gi...@apache.org>.
RussellSpitzer edited a comment on issue #1894:
URL: https://github.com/apache/iceberg/issues/1894#issuecomment-774276183
Already Close File tends to be that the data was not actually partitioned on the correct columns. This is an error that says the data within a Spark Partition contained data for multiple Iceberg Partitions and that this data was not ordered correctly. IE there was a record for Iceberg partition A, then one for Iceberg partition B, then one for A again.
So my guess would be that `partition_by_columns` does not match the table
----------------------------------------------------------------
This is an automated message from the Apache Git Service.
To respond to the message, please log on to GitHub and use the
URL above to go to the specific comment.
For queries about this service, please contact Infrastructure at:
users@infra.apache.org
---------------------------------------------------------------------
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For additional commands, e-mail: issues-help@iceberg.apache.org
[GitHub] [iceberg] RussellSpitzer edited a comment on issue #1894: Spark writes the Iceberg dual partition table to report an error
Posted by GitBox <gi...@apache.org>.
RussellSpitzer edited a comment on issue #1894:
URL: https://github.com/apache/iceberg/issues/1894#issuecomment-774303754
All Iceberg tables exist in the context of a Catalog. If you haven't specified one and just provided a path it is using the implied "HadoopCatalog". The table will have It's own Metadata.json file which will specify the schema of the table and partition spec. Once created, that is the schema and partition spec until it is specifically altered.
So in your example "./data-iceberg" will be treated as a HadoopTable and look in that directory for a Metadata.json with a specified schema and partitioning. If such a file did not exist you would get an error like
```scala
spark.read.parquet("/Users/russellspitzer/Temp/parquet_data").write.format("iceberg").save("/Users/russellspitzer/Temp/iceberg_table")
org.apache.iceberg.exceptions.NoSuchTableException: Table does not exist at location: /Users/russellspitzer/Temp/iceberg_table
```
Since you didn't get such an error means that a table does exist there and it has a schema.
When attempting attempting to write Spark saw "partitionBy" "a", "b", and preformed a shuffle on the input data
So all spark partitions are now like
```
SparkPartition 1 - > (1,1,1,1) , (1,1,2,1), (1,1,1,3) ...
SparkPartition 2 -> (1,2,1,1), (1,2,2,1), (1,2,1,2) ...
SparkPartition 3 -> (2,4,1,1), (2,4,2,1), (2,4,1,2) ...
```
The first two values are always the same within the spark partition, but the last two are in a random order.
Then the Iceberg writer gets one of these partitions and it creates a writer based on it's Metadata.json which let's say is not
the same as the `partitionedBy` argument. For this example let's say the table is actually partitioned by 'c'
By default in Spark Iceberg's writer opens a file for each partition assumes it will write all the values for that partition, then close the file. If it tries to write partition 1 it ends up doing the following
```
(1,1,1,1) -> Open partition c = 1 and start new file
(1,1,2,1) -> Close partition c = 1 file; Open partition c =2 and start new file
(1,1,1,3) -> Open partition c = 1 /// Error already closed file for Partition c=1
```
Since this is a runtime check you won't see this error with a single row since that will always be able to open and close the file without incident, but a dataset with multiple rows can potentially trigger this. Most of the time you want this Spark partitioning to match the Iceberg partitioning because this minimizes the number of files you need to actually write.
If you really don't want to have the Spark Partitioning match for some reason you can use the sortWithinPartitions to order the data within the partitions by their Iceberg partition values.
The other possibility is to use the new spark fanout writer which keeps all of the files open until they entire task has been completed.
Neither of these solutions is really as good as having your data partitioned like the data is expecting since that minimizes the number of files being written since all the files for a given partition will be done by the same Spark Task.
----------------------------------------------------------------
This is an automated message from the Apache Git Service.
To respond to the message, please log on to GitHub and use the
URL above to go to the specific comment.
For queries about this service, please contact Infrastructure at:
users@infra.apache.org
---------------------------------------------------------------------
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For additional commands, e-mail: issues-help@iceberg.apache.org
[GitHub] [iceberg] Ranjith-AR commented on issue #1894: Spark writes the Iceberg dual partition table to report an error
Posted by GitBox <gi...@apache.org>.
Ranjith-AR commented on issue #1894:
URL: https://github.com/apache/iceberg/issues/1894#issuecomment-1050738332
I tried replicating the dual partition example provided in the docs
https://iceberg.apache.org/docs/latest/spark-writes/

Created the below table:
-------
"CREATE TABLE nyc.green_taxis_by_days_vendorid (
VendorID string,
lpep_pickup_datetime timestamp,
lpep_dropoff_datetime string,
store_and_fwd_flag string,RatecodeID string,PULocationID string,DOLocationID string,
passenger_count string,trip_distance string,fare_amount string,extra string,mta_tax string,
tip_amount string,tolls_amount string,ehail_fee string,improvement_surcharge string,
total_amount string,
payment_type string,trip_type string,
congestion_surcharge string)
USING iceberg
PARTITIONED BY (
days(lpep_pickup_datetime),VendorID
)"
-----
Trying to write into the table via the dataframe api.
df_green_stage1.sortWithinPartitions(("lpep_pickup_datetime"),"VendorID") \
.writeTo("nyc.green_taxis_by_days_vendorid") \
.append()
------
However, I run into the below error
The partitions are defined as days(timestamp) and secondary as vendor id.
since the dataframe is sorted by "timestamp, vendor id" and not as "days(timestamp), vendor id" as defined in the iceberg table , we get the file already closed error.
Is this a bug ? any work arounds (other than adding an additional date column and using that in the sort) ?
--------
An error was encountered:
An error occurred while calling o294.append.
: org.apache.spark.SparkException: Writing job aborted.
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2(WriteToDataSourceV2Exec.scala:388)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2$(WriteToDataSourceV2Exec.scala:336)
at org.apache.spark.sql.execution.datasources.v2.AppendDataExec.writeWithV2(WriteToDataSourceV2Exec.scala:218)
at org.apache.spark.sql.execution.datasources.v2.AppendDataExec.run(WriteToDataSourceV2Exec.scala:225)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.result$lzycompute(V2CommandExec.scala:40)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.result(V2CommandExec.scala:40)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.doExecute(V2CommandExec.scala:55)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$execute$1(SparkPlan.scala:194)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$executeQuery$1(SparkPlan.scala:232)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:151)
at org.apache.spark.sql.execution.SparkPlan.executeQuery(SparkPlan.scala:229)
at org.apache.spark.sql.execution.SparkPlan.execute(SparkPlan.scala:190)
at org.apache.spark.sql.execution.QueryExecution.toRdd$lzycompute(QueryExecution.scala:134)
at org.apache.spark.sql.execution.QueryExecution.toRdd(QueryExecution.scala:133)
at org.apache.spark.sql.DataFrameWriterV2.$anonfun$runCommand$1(DataFrameWriterV2.scala:196)
at org.apache.spark.sql.catalyst.QueryPlanningTracker$.withTracker(QueryPlanningTracker.scala:107)
at org.apache.spark.sql.execution.SQLExecution$.withTracker(SQLExecution.scala:232)
at org.apache.spark.sql.execution.SQLExecution$.executeQuery$1(SQLExecution.scala:110)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$6(SQLExecution.scala:135)
at org.apache.spark.sql.catalyst.QueryPlanningTracker$.withTracker(QueryPlanningTracker.scala:107)
at org.apache.spark.sql.execution.SQLExecution$.withTracker(SQLExecution.scala:232)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$5(SQLExecution.scala:135)
at org.apache.spark.sql.execution.SQLExecution$.withSQLConfPropagated(SQLExecution.scala:253)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$1(SQLExecution.scala:134)
at org.apache.spark.sql.SparkSession.withActive(SparkSession.scala:775)
at org.apache.spark.sql.execution.SQLExecution$.withNewExecutionId(SQLExecution.scala:68)
at org.apache.spark.sql.DataFrameWriterV2.runCommand(DataFrameWriterV2.scala:196)
at org.apache.spark.sql.DataFrameWriterV2.append(DataFrameWriterV2.scala:149)
at sun.reflect.NativeMethodAccessorImpl.invoke0(Native Method)
at sun.reflect.NativeMethodAccessorImpl.invoke(NativeMethodAccessorImpl.java:62)
at sun.reflect.DelegatingMethodAccessorImpl.invoke(DelegatingMethodAccessorImpl.java:43)
at java.lang.reflect.Method.invoke(Method.java:498)
at py4j.reflection.MethodInvoker.invoke(MethodInvoker.java:244)
at py4j.reflection.ReflectionEngine.invoke(ReflectionEngine.java:357)
at py4j.Gateway.invoke(Gateway.java:282)
at py4j.commands.AbstractCommand.invokeMethod(AbstractCommand.java:132)
at py4j.commands.CallCommand.execute(CallCommand.java:79)
at py4j.GatewayConnection.run(GatewayConnection.java:238)
at java.lang.Thread.run(Thread.java:750)
Caused by: org.apache.spark.SparkException: Job aborted due to stage failure: Task 0 in stage 32.0 failed 4 times, most recent failure: Lost task 0.3 in stage 32.0 (TID 103) (ip-172-31-21-243.ec2.internal executor 18): java.lang.IllegalStateException: Incoming records violate the writer assumption that records are clustered by spec and by partition within each spec. Either cluster the incoming records or switch to fanout writers.
Encountered records that belong to already closed files:
partition 'lpep_pickup_datetime_day=2020-04-01/VendorID=2' in spec [
1000: lpep_pickup_datetime_day: day(2)
1001: VendorID: identity(1)
]
at org.apache.iceberg.io.ClusteredWriter.write(ClusteredWriter.java:95)
at org.apache.iceberg.io.ClusteredDataWriter.write(ClusteredDataWriter.java:34)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:629)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:604)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.$anonfun$run$1(WriteToDataSourceV2Exec.scala:416)
at org.apache.spark.util.Utils$.tryWithSafeFinallyAndFailureCallbacks(Utils.scala:1473)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.run(WriteToDataSourceV2Exec.scala:452)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.$anonfun$writeWithV2$2(WriteToDataSourceV2Exec.scala:360)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:131)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:497)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1439)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:500)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
at java.lang.Thread.run(Thread.java:750)
Driver stacktrace:
at org.apache.spark.scheduler.DAGScheduler.failJobAndIndependentStages(DAGScheduler.scala:2470)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2(DAGScheduler.scala:2419)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2$adapted(DAGScheduler.scala:2418)
at scala.collection.mutable.ResizableArray.foreach(ResizableArray.scala:62)
at scala.collection.mutable.ResizableArray.foreach$(ResizableArray.scala:55)
at scala.collection.mutable.ArrayBuffer.foreach(ArrayBuffer.scala:49)
at org.apache.spark.scheduler.DAGScheduler.abortStage(DAGScheduler.scala:2418)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1(DAGScheduler.scala:1125)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1$adapted(DAGScheduler.scala:1125)
at scala.Option.foreach(Option.scala:407)
at org.apache.spark.scheduler.DAGScheduler.handleTaskSetFailed(DAGScheduler.scala:1125)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.doOnReceive(DAGScheduler.scala:2684)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2626)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2615)
at org.apache.spark.util.EventLoop$$anon$1.run(EventLoop.scala:49)
at org.apache.spark.scheduler.DAGScheduler.runJob(DAGScheduler.scala:914)
at org.apache.spark.SparkContext.runJob(SparkContext.scala:2241)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2(WriteToDataSourceV2Exec.scala:357)
... 38 more
Caused by: java.lang.IllegalStateException: Incoming records violate the writer assumption that records are clustered by spec and by partition within each spec. Either cluster the incoming records or switch to fanout writers.
Encountered records that belong to already closed files:
partition 'lpep_pickup_datetime_day=2020-04-01/VendorID=2' in spec [
1000: lpep_pickup_datetime_day: day(2)
1001: VendorID: identity(1)
]
at org.apache.iceberg.io.ClusteredWriter.write(ClusteredWriter.java:95)
at org.apache.iceberg.io.ClusteredDataWriter.write(ClusteredDataWriter.java:34)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:629)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:604)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.$anonfun$run$1(WriteToDataSourceV2Exec.scala:416)
at org.apache.spark.util.Utils$.tryWithSafeFinallyAndFailureCallbacks(Utils.scala:1473)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.run(WriteToDataSourceV2Exec.scala:452)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.$anonfun$writeWithV2$2(WriteToDataSourceV2Exec.scala:360)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:131)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:497)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1439)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:500)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
... 1 more
Traceback (most recent call last):
File "/usr/lib/spark/python/lib/pyspark.zip/pyspark/sql/readwriter.py", line 1564, in append
self._jwriter.append()
File "/usr/lib/spark/python/lib/py4j-0.10.9-src.zip/py4j/java_gateway.py", line 1305, in __call__
answer, self.gateway_client, self.target_id, self.name)
File "/usr/lib/spark/python/lib/pyspark.zip/pyspark/sql/utils.py", line 111, in deco
return f(*a, **kw)
File "/usr/lib/spark/python/lib/py4j-0.10.9-src.zip/py4j/protocol.py", line 328, in get_return_value
format(target_id, ".", name), value)
py4j.protocol.Py4JJavaError: An error occurred while calling o294.append.
: org.apache.spark.SparkException: Writing job aborted.
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2(WriteToDataSourceV2Exec.scala:388)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2$(WriteToDataSourceV2Exec.scala:336)
at org.apache.spark.sql.execution.datasources.v2.AppendDataExec.writeWithV2(WriteToDataSourceV2Exec.scala:218)
at org.apache.spark.sql.execution.datasources.v2.AppendDataExec.run(WriteToDataSourceV2Exec.scala:225)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.result$lzycompute(V2CommandExec.scala:40)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.result(V2CommandExec.scala:40)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.doExecute(V2CommandExec.scala:55)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$execute$1(SparkPlan.scala:194)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$executeQuery$1(SparkPlan.scala:232)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:151)
at org.apache.spark.sql.execution.SparkPlan.executeQuery(SparkPlan.scala:229)
at org.apache.spark.sql.execution.SparkPlan.execute(SparkPlan.scala:190)
at org.apache.spark.sql.execution.QueryExecution.toRdd$lzycompute(QueryExecution.scala:134)
at org.apache.spark.sql.execution.QueryExecution.toRdd(QueryExecution.scala:133)
at org.apache.spark.sql.DataFrameWriterV2.$anonfun$runCommand$1(DataFrameWriterV2.scala:196)
at org.apache.spark.sql.catalyst.QueryPlanningTracker$.withTracker(QueryPlanningTracker.scala:107)
at org.apache.spark.sql.execution.SQLExecution$.withTracker(SQLExecution.scala:232)
at org.apache.spark.sql.execution.SQLExecution$.executeQuery$1(SQLExecution.scala:110)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$6(SQLExecution.scala:135)
at org.apache.spark.sql.catalyst.QueryPlanningTracker$.withTracker(QueryPlanningTracker.scala:107)
at org.apache.spark.sql.execution.SQLExecution$.withTracker(SQLExecution.scala:232)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$5(SQLExecution.scala:135)
at org.apache.spark.sql.execution.SQLExecution$.withSQLConfPropagated(SQLExecution.scala:253)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$1(SQLExecution.scala:134)
at org.apache.spark.sql.SparkSession.withActive(SparkSession.scala:775)
at org.apache.spark.sql.execution.SQLExecution$.withNewExecutionId(SQLExecution.scala:68)
at org.apache.spark.sql.DataFrameWriterV2.runCommand(DataFrameWriterV2.scala:196)
at org.apache.spark.sql.DataFrameWriterV2.append(DataFrameWriterV2.scala:149)
at sun.reflect.NativeMethodAccessorImpl.invoke0(Native Method)
at sun.reflect.NativeMethodAccessorImpl.invoke(NativeMethodAccessorImpl.java:62)
at sun.reflect.DelegatingMethodAccessorImpl.invoke(DelegatingMethodAccessorImpl.java:43)
at java.lang.reflect.Method.invoke(Method.java:498)
at py4j.reflection.MethodInvoker.invoke(MethodInvoker.java:244)
at py4j.reflection.ReflectionEngine.invoke(ReflectionEngine.java:357)
at py4j.Gateway.invoke(Gateway.java:282)
at py4j.commands.AbstractCommand.invokeMethod(AbstractCommand.java:132)
at py4j.commands.CallCommand.execute(CallCommand.java:79)
at py4j.GatewayConnection.run(GatewayConnection.java:238)
at java.lang.Thread.run(Thread.java:750)
Caused by: org.apache.spark.SparkException: Job aborted due to stage failure: Task 0 in stage 32.0 failed 4 times, most recent failure: Lost task 0.3 in stage 32.0 (TID 103) (ip-172-31-21-243.ec2.internal executor 18): java.lang.IllegalStateException: Incoming records violate the writer assumption that records are clustered by spec and by partition within each spec. Either cluster the incoming records or switch to fanout writers.
Encountered records that belong to already closed files:
partition 'lpep_pickup_datetime_day=2020-04-01/VendorID=2' in spec [
1000: lpep_pickup_datetime_day: day(2)
1001: VendorID: identity(1)
]
at org.apache.iceberg.io.ClusteredWriter.write(ClusteredWriter.java:95)
at org.apache.iceberg.io.ClusteredDataWriter.write(ClusteredDataWriter.java:34)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:629)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:604)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.$anonfun$run$1(WriteToDataSourceV2Exec.scala:416)
at org.apache.spark.util.Utils$.tryWithSafeFinallyAndFailureCallbacks(Utils.scala:1473)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.run(WriteToDataSourceV2Exec.scala:452)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.$anonfun$writeWithV2$2(WriteToDataSourceV2Exec.scala:360)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:131)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:497)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1439)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:500)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
at java.lang.Thread.run(Thread.java:750)
Driver stacktrace:
at org.apache.spark.scheduler.DAGScheduler.failJobAndIndependentStages(DAGScheduler.scala:2470)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2(DAGScheduler.scala:2419)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2$adapted(DAGScheduler.scala:2418)
at scala.collection.mutable.ResizableArray.foreach(ResizableArray.scala:62)
at scala.collection.mutable.ResizableArray.foreach$(ResizableArray.scala:55)
at scala.collection.mutable.ArrayBuffer.foreach(ArrayBuffer.scala:49)
at org.apache.spark.scheduler.DAGScheduler.abortStage(DAGScheduler.scala:2418)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1(DAGScheduler.scala:1125)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1$adapted(DAGScheduler.scala:1125)
at scala.Option.foreach(Option.scala:407)
at org.apache.spark.scheduler.DAGScheduler.handleTaskSetFailed(DAGScheduler.scala:1125)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.doOnReceive(DAGScheduler.scala:2684)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2626)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2615)
at org.apache.spark.util.EventLoop$$anon$1.run(EventLoop.scala:49)
at org.apache.spark.scheduler.DAGScheduler.runJob(DAGScheduler.scala:914)
at org.apache.spark.SparkContext.runJob(SparkContext.scala:2241)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2(WriteToDataSourceV2Exec.scala:357)
... 38 more
Caused by: java.lang.IllegalStateException: Incoming records violate the writer assumption that records are clustered by spec and by partition within each spec. Either cluster the incoming records or switch to fanout writers.
Encountered records that belong to already closed files:
partition 'lpep_pickup_datetime_day=2020-04-01/VendorID=2' in spec [
1000: lpep_pickup_datetime_day: day(2)
1001: VendorID: identity(1)
]
at org.apache.iceberg.io.ClusteredWriter.write(ClusteredWriter.java:95)
at org.apache.iceberg.io.ClusteredDataWriter.write(ClusteredDataWriter.java:34)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:629)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:604)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.$anonfun$run$1(WriteToDataSourceV2Exec.scala:416)
at org.apache.spark.util.Utils$.tryWithSafeFinallyAndFailureCallbacks(Utils.scala:1473)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.run(WriteToDataSourceV2Exec.scala:452)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.$anonfun$writeWithV2$2(WriteToDataSourceV2Exec.scala:360)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:131)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:497)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1439)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:500)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
... 1 more
-------
--
This is an automated message from the Apache Git Service.
To respond to the message, please log on to GitHub and use the
URL above to go to the specific comment.
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For queries about this service, please contact Infrastructure at:
users@infra.apache.org
---------------------------------------------------------------------
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For additional commands, e-mail: issues-help@iceberg.apache.org
[GitHub] [iceberg] RussellSpitzer commented on issue #1894: Spark writes the Iceberg dual partition table to report an error
Posted by GitBox <gi...@apache.org>.
RussellSpitzer commented on issue #1894:
URL: https://github.com/apache/iceberg/issues/1894#issuecomment-774305299
https://iceberg.apache.org/spark-writes/#writing-to-partitioned-tables - Docs on this issue
----------------------------------------------------------------
This is an automated message from the Apache Git Service.
To respond to the message, please log on to GitHub and use the
URL above to go to the specific comment.
For queries about this service, please contact Infrastructure at:
users@infra.apache.org
---------------------------------------------------------------------
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For additional commands, e-mail: issues-help@iceberg.apache.org
[GitHub] [iceberg] holdenk commented on issue #1894: Spark writes the Iceberg dual partition table to report an error
Posted by GitBox <gi...@apache.org>.
holdenk commented on issue #1894:
URL: https://github.com/apache/iceberg/issues/1894#issuecomment-749164004
Do you have a full stack trace?
----------------------------------------------------------------
This is an automated message from the Apache Git Service.
To respond to the message, please log on to GitHub and use the
URL above to go to the specific comment.
For queries about this service, please contact Infrastructure at:
users@infra.apache.org
---------------------------------------------------------------------
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For additional commands, e-mail: issues-help@iceberg.apache.org
[GitHub] [iceberg] RussellSpitzer removed a comment on issue #1894: Spark writes the Iceberg dual partition table to report an error
Posted by GitBox <gi...@apache.org>.
RussellSpitzer removed a comment on issue #1894:
URL: https://github.com/apache/iceberg/issues/1894#issuecomment-774307653
And here is an example
```scala
scala> spark.sql("CREATE table ice.default.part (x Int, y Int) USING iceberg partitioned by (x) location '/Users/russellspitzer/Temp/iceberg_table'");
val df = spark.createDataFrame(Seq((1,1),(1,2),(1,3),(2,1),(2,2),(2,3),(3,1),(3,2),(3,3))).toDF("x","y")
scala> df.write.format("iceberg").partitionBy("x").mode("overwrite").saveAsTable("ice.default.part")
// No Error
scala> df.write.format("iceberg").partitionBy("y").mode("overwrite").saveAsTable("ice.default.part")
// Many errors
// Caused by: java.lang.IllegalStateException: Already closed files for partition: y=1
```
----------------------------------------------------------------
This is an automated message from the Apache Git Service.
To respond to the message, please log on to GitHub and use the
URL above to go to the specific comment.
For queries about this service, please contact Infrastructure at:
users@infra.apache.org
---------------------------------------------------------------------
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For additional commands, e-mail: issues-help@iceberg.apache.org
[GitHub] [iceberg] Ranjith-AR removed a comment on issue #1894: Spark writes the Iceberg dual partition table to report an error
Posted by GitBox <gi...@apache.org>.
Ranjith-AR removed a comment on issue #1894:
URL: https://github.com/apache/iceberg/issues/1894#issuecomment-1050738332
I tried replicating the dual partition example provided in the docs
https://iceberg.apache.org/docs/latest/spark-writes/

Created the below table:
-------
"CREATE TABLE nyc.green_taxis_by_days_vendorid (
VendorID string,
lpep_pickup_datetime timestamp,
lpep_dropoff_datetime string,
store_and_fwd_flag string,RatecodeID string,PULocationID string,DOLocationID string,
passenger_count string,trip_distance string,fare_amount string,extra string,mta_tax string,
tip_amount string,tolls_amount string,ehail_fee string,improvement_surcharge string,
total_amount string,
payment_type string,trip_type string,
congestion_surcharge string)
USING iceberg
PARTITIONED BY (
***days(lpep_pickup_datetime),VendorID***
)"
-----
Trying to write into the table via both dataframe api and pyspark sql...and get the same error (i.e. file already closed)
df_green_stage1.sortWithinPartitions(("lpep_pickup_datetime"),"VendorID") \
.writeTo("nyc.green_taxis_by_days_vendorid") \
.append()
-----
%%sql
INSERT INTO nyc.green_taxis_by_days_vendorid (
SELECT
VendorID,lpep_pickup_datetime,lpep_dropoff_datetime,store_and_fwd_flag,RatecodeID,PULocationID,DOLocationID,passenger_count,trip_distance,fare_amount,
extra,mta_tax,tip_amount,tolls_amount,ehail_fee,
improvement_surcharge,total_amount,payment_type,trip_type,congestion_surcharge
FROM nyc.green_taxis_by_days_vendorid_withdate
ORDER BY lpep_pickup_datetime,VendorID
)
------
However, I run into the below error
The partitions are defined as days(timestamp) and secondary as vendor id.
since the dataframe is sorted by "timestamp, vendor id" and not as "days(timestamp), vendor id" as defined in the iceberg table , we get the file already closed error.

Is this a bug ? any work arounds (other than adding an additional date column and using that in the sort) ?
--------
An error was encountered:
An error occurred while calling o294.append.
: org.apache.spark.SparkException: Writing job aborted.
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2(WriteToDataSourceV2Exec.scala:388)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2$(WriteToDataSourceV2Exec.scala:336)
at org.apache.spark.sql.execution.datasources.v2.AppendDataExec.writeWithV2(WriteToDataSourceV2Exec.scala:218)
at org.apache.spark.sql.execution.datasources.v2.AppendDataExec.run(WriteToDataSourceV2Exec.scala:225)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.result$lzycompute(V2CommandExec.scala:40)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.result(V2CommandExec.scala:40)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.doExecute(V2CommandExec.scala:55)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$execute$1(SparkPlan.scala:194)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$executeQuery$1(SparkPlan.scala:232)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:151)
at org.apache.spark.sql.execution.SparkPlan.executeQuery(SparkPlan.scala:229)
at org.apache.spark.sql.execution.SparkPlan.execute(SparkPlan.scala:190)
at org.apache.spark.sql.execution.QueryExecution.toRdd$lzycompute(QueryExecution.scala:134)
at org.apache.spark.sql.execution.QueryExecution.toRdd(QueryExecution.scala:133)
at org.apache.spark.sql.DataFrameWriterV2.$anonfun$runCommand$1(DataFrameWriterV2.scala:196)
at org.apache.spark.sql.catalyst.QueryPlanningTracker$.withTracker(QueryPlanningTracker.scala:107)
at org.apache.spark.sql.execution.SQLExecution$.withTracker(SQLExecution.scala:232)
at org.apache.spark.sql.execution.SQLExecution$.executeQuery$1(SQLExecution.scala:110)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$6(SQLExecution.scala:135)
at org.apache.spark.sql.catalyst.QueryPlanningTracker$.withTracker(QueryPlanningTracker.scala:107)
at org.apache.spark.sql.execution.SQLExecution$.withTracker(SQLExecution.scala:232)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$5(SQLExecution.scala:135)
at org.apache.spark.sql.execution.SQLExecution$.withSQLConfPropagated(SQLExecution.scala:253)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$1(SQLExecution.scala:134)
at org.apache.spark.sql.SparkSession.withActive(SparkSession.scala:775)
at org.apache.spark.sql.execution.SQLExecution$.withNewExecutionId(SQLExecution.scala:68)
at org.apache.spark.sql.DataFrameWriterV2.runCommand(DataFrameWriterV2.scala:196)
at org.apache.spark.sql.DataFrameWriterV2.append(DataFrameWriterV2.scala:149)
at sun.reflect.NativeMethodAccessorImpl.invoke0(Native Method)
at sun.reflect.NativeMethodAccessorImpl.invoke(NativeMethodAccessorImpl.java:62)
at sun.reflect.DelegatingMethodAccessorImpl.invoke(DelegatingMethodAccessorImpl.java:43)
at java.lang.reflect.Method.invoke(Method.java:498)
at py4j.reflection.MethodInvoker.invoke(MethodInvoker.java:244)
at py4j.reflection.ReflectionEngine.invoke(ReflectionEngine.java:357)
at py4j.Gateway.invoke(Gateway.java:282)
at py4j.commands.AbstractCommand.invokeMethod(AbstractCommand.java:132)
at py4j.commands.CallCommand.execute(CallCommand.java:79)
at py4j.GatewayConnection.run(GatewayConnection.java:238)
at java.lang.Thread.run(Thread.java:750)
Caused by: org.apache.spark.SparkException: Job aborted due to stage failure: Task 0 in stage 32.0 failed 4 times, most recent failure: Lost task 0.3 in stage 32.0 (TID 103) (ip-172-31-21-243.ec2.internal executor 18): java.lang.IllegalStateException: Incoming records violate the writer assumption that records are clustered by spec and by partition within each spec. Either cluster the incoming records or switch to fanout writers.
Encountered records that belong to already closed files:
partition 'lpep_pickup_datetime_day=2020-04-01/VendorID=2' in spec [
1000: lpep_pickup_datetime_day: day(2)
1001: VendorID: identity(1)
]
at org.apache.iceberg.io.ClusteredWriter.write(ClusteredWriter.java:95)
at org.apache.iceberg.io.ClusteredDataWriter.write(ClusteredDataWriter.java:34)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:629)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:604)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.$anonfun$run$1(WriteToDataSourceV2Exec.scala:416)
at org.apache.spark.util.Utils$.tryWithSafeFinallyAndFailureCallbacks(Utils.scala:1473)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.run(WriteToDataSourceV2Exec.scala:452)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.$anonfun$writeWithV2$2(WriteToDataSourceV2Exec.scala:360)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:131)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:497)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1439)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:500)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
at java.lang.Thread.run(Thread.java:750)
Driver stacktrace:
at org.apache.spark.scheduler.DAGScheduler.failJobAndIndependentStages(DAGScheduler.scala:2470)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2(DAGScheduler.scala:2419)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2$adapted(DAGScheduler.scala:2418)
at scala.collection.mutable.ResizableArray.foreach(ResizableArray.scala:62)
at scala.collection.mutable.ResizableArray.foreach$(ResizableArray.scala:55)
at scala.collection.mutable.ArrayBuffer.foreach(ArrayBuffer.scala:49)
at org.apache.spark.scheduler.DAGScheduler.abortStage(DAGScheduler.scala:2418)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1(DAGScheduler.scala:1125)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1$adapted(DAGScheduler.scala:1125)
at scala.Option.foreach(Option.scala:407)
at org.apache.spark.scheduler.DAGScheduler.handleTaskSetFailed(DAGScheduler.scala:1125)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.doOnReceive(DAGScheduler.scala:2684)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2626)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2615)
at org.apache.spark.util.EventLoop$$anon$1.run(EventLoop.scala:49)
at org.apache.spark.scheduler.DAGScheduler.runJob(DAGScheduler.scala:914)
at org.apache.spark.SparkContext.runJob(SparkContext.scala:2241)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2(WriteToDataSourceV2Exec.scala:357)
... 38 more
Caused by: java.lang.IllegalStateException: Incoming records violate the writer assumption that records are clustered by spec and by partition within each spec. Either cluster the incoming records or switch to fanout writers.
Encountered records that belong to already closed files:
partition 'lpep_pickup_datetime_day=2020-04-01/VendorID=2' in spec [
1000: lpep_pickup_datetime_day: day(2)
1001: VendorID: identity(1)
]
at org.apache.iceberg.io.ClusteredWriter.write(ClusteredWriter.java:95)
at org.apache.iceberg.io.ClusteredDataWriter.write(ClusteredDataWriter.java:34)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:629)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:604)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.$anonfun$run$1(WriteToDataSourceV2Exec.scala:416)
at org.apache.spark.util.Utils$.tryWithSafeFinallyAndFailureCallbacks(Utils.scala:1473)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.run(WriteToDataSourceV2Exec.scala:452)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.$anonfun$writeWithV2$2(WriteToDataSourceV2Exec.scala:360)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:131)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:497)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1439)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:500)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
... 1 more
Traceback (most recent call last):
File "/usr/lib/spark/python/lib/pyspark.zip/pyspark/sql/readwriter.py", line 1564, in append
self._jwriter.append()
File "/usr/lib/spark/python/lib/py4j-0.10.9-src.zip/py4j/java_gateway.py", line 1305, in __call__
answer, self.gateway_client, self.target_id, self.name)
File "/usr/lib/spark/python/lib/pyspark.zip/pyspark/sql/utils.py", line 111, in deco
return f(*a, **kw)
File "/usr/lib/spark/python/lib/py4j-0.10.9-src.zip/py4j/protocol.py", line 328, in get_return_value
format(target_id, ".", name), value)
py4j.protocol.Py4JJavaError: An error occurred while calling o294.append.
: org.apache.spark.SparkException: Writing job aborted.
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2(WriteToDataSourceV2Exec.scala:388)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2$(WriteToDataSourceV2Exec.scala:336)
at org.apache.spark.sql.execution.datasources.v2.AppendDataExec.writeWithV2(WriteToDataSourceV2Exec.scala:218)
at org.apache.spark.sql.execution.datasources.v2.AppendDataExec.run(WriteToDataSourceV2Exec.scala:225)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.result$lzycompute(V2CommandExec.scala:40)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.result(V2CommandExec.scala:40)
at org.apache.spark.sql.execution.datasources.v2.V2CommandExec.doExecute(V2CommandExec.scala:55)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$execute$1(SparkPlan.scala:194)
at org.apache.spark.sql.execution.SparkPlan.$anonfun$executeQuery$1(SparkPlan.scala:232)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:151)
at org.apache.spark.sql.execution.SparkPlan.executeQuery(SparkPlan.scala:229)
at org.apache.spark.sql.execution.SparkPlan.execute(SparkPlan.scala:190)
at org.apache.spark.sql.execution.QueryExecution.toRdd$lzycompute(QueryExecution.scala:134)
at org.apache.spark.sql.execution.QueryExecution.toRdd(QueryExecution.scala:133)
at org.apache.spark.sql.DataFrameWriterV2.$anonfun$runCommand$1(DataFrameWriterV2.scala:196)
at org.apache.spark.sql.catalyst.QueryPlanningTracker$.withTracker(QueryPlanningTracker.scala:107)
at org.apache.spark.sql.execution.SQLExecution$.withTracker(SQLExecution.scala:232)
at org.apache.spark.sql.execution.SQLExecution$.executeQuery$1(SQLExecution.scala:110)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$6(SQLExecution.scala:135)
at org.apache.spark.sql.catalyst.QueryPlanningTracker$.withTracker(QueryPlanningTracker.scala:107)
at org.apache.spark.sql.execution.SQLExecution$.withTracker(SQLExecution.scala:232)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$5(SQLExecution.scala:135)
at org.apache.spark.sql.execution.SQLExecution$.withSQLConfPropagated(SQLExecution.scala:253)
at org.apache.spark.sql.execution.SQLExecution$.$anonfun$withNewExecutionId$1(SQLExecution.scala:134)
at org.apache.spark.sql.SparkSession.withActive(SparkSession.scala:775)
at org.apache.spark.sql.execution.SQLExecution$.withNewExecutionId(SQLExecution.scala:68)
at org.apache.spark.sql.DataFrameWriterV2.runCommand(DataFrameWriterV2.scala:196)
at org.apache.spark.sql.DataFrameWriterV2.append(DataFrameWriterV2.scala:149)
at sun.reflect.NativeMethodAccessorImpl.invoke0(Native Method)
at sun.reflect.NativeMethodAccessorImpl.invoke(NativeMethodAccessorImpl.java:62)
at sun.reflect.DelegatingMethodAccessorImpl.invoke(DelegatingMethodAccessorImpl.java:43)
at java.lang.reflect.Method.invoke(Method.java:498)
at py4j.reflection.MethodInvoker.invoke(MethodInvoker.java:244)
at py4j.reflection.ReflectionEngine.invoke(ReflectionEngine.java:357)
at py4j.Gateway.invoke(Gateway.java:282)
at py4j.commands.AbstractCommand.invokeMethod(AbstractCommand.java:132)
at py4j.commands.CallCommand.execute(CallCommand.java:79)
at py4j.GatewayConnection.run(GatewayConnection.java:238)
at java.lang.Thread.run(Thread.java:750)
Caused by: org.apache.spark.SparkException: Job aborted due to stage failure: Task 0 in stage 32.0 failed 4 times, most recent failure: Lost task 0.3 in stage 32.0 (TID 103) (ip-172-31-21-243.ec2.internal executor 18): java.lang.IllegalStateException: Incoming records violate the writer assumption that records are clustered by spec and by partition within each spec. Either cluster the incoming records or switch to fanout writers.
Encountered records that belong to already closed files:
partition 'lpep_pickup_datetime_day=2020-04-01/VendorID=2' in spec [
1000: lpep_pickup_datetime_day: day(2)
1001: VendorID: identity(1)
]
at org.apache.iceberg.io.ClusteredWriter.write(ClusteredWriter.java:95)
at org.apache.iceberg.io.ClusteredDataWriter.write(ClusteredDataWriter.java:34)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:629)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:604)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.$anonfun$run$1(WriteToDataSourceV2Exec.scala:416)
at org.apache.spark.util.Utils$.tryWithSafeFinallyAndFailureCallbacks(Utils.scala:1473)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.run(WriteToDataSourceV2Exec.scala:452)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.$anonfun$writeWithV2$2(WriteToDataSourceV2Exec.scala:360)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:131)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:497)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1439)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:500)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
at java.lang.Thread.run(Thread.java:750)
Driver stacktrace:
at org.apache.spark.scheduler.DAGScheduler.failJobAndIndependentStages(DAGScheduler.scala:2470)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2(DAGScheduler.scala:2419)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$abortStage$2$adapted(DAGScheduler.scala:2418)
at scala.collection.mutable.ResizableArray.foreach(ResizableArray.scala:62)
at scala.collection.mutable.ResizableArray.foreach$(ResizableArray.scala:55)
at scala.collection.mutable.ArrayBuffer.foreach(ArrayBuffer.scala:49)
at org.apache.spark.scheduler.DAGScheduler.abortStage(DAGScheduler.scala:2418)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1(DAGScheduler.scala:1125)
at org.apache.spark.scheduler.DAGScheduler.$anonfun$handleTaskSetFailed$1$adapted(DAGScheduler.scala:1125)
at scala.Option.foreach(Option.scala:407)
at org.apache.spark.scheduler.DAGScheduler.handleTaskSetFailed(DAGScheduler.scala:1125)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.doOnReceive(DAGScheduler.scala:2684)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2626)
at org.apache.spark.scheduler.DAGSchedulerEventProcessLoop.onReceive(DAGScheduler.scala:2615)
at org.apache.spark.util.EventLoop$$anon$1.run(EventLoop.scala:49)
at org.apache.spark.scheduler.DAGScheduler.runJob(DAGScheduler.scala:914)
at org.apache.spark.SparkContext.runJob(SparkContext.scala:2241)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.writeWithV2(WriteToDataSourceV2Exec.scala:357)
... 38 more
Caused by: java.lang.IllegalStateException: Incoming records violate the writer assumption that records are clustered by spec and by partition within each spec. Either cluster the incoming records or switch to fanout writers.
Encountered records that belong to already closed files:
partition 'lpep_pickup_datetime_day=2020-04-01/VendorID=2' in spec [
1000: lpep_pickup_datetime_day: day(2)
1001: VendorID: identity(1)
]
at org.apache.iceberg.io.ClusteredWriter.write(ClusteredWriter.java:95)
at org.apache.iceberg.io.ClusteredDataWriter.write(ClusteredDataWriter.java:34)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:629)
at org.apache.iceberg.spark.source.SparkWrite$PartitionedDataWriter.write(SparkWrite.java:604)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.$anonfun$run$1(WriteToDataSourceV2Exec.scala:416)
at org.apache.spark.util.Utils$.tryWithSafeFinallyAndFailureCallbacks(Utils.scala:1473)
at org.apache.spark.sql.execution.datasources.v2.DataWritingSparkTask$.run(WriteToDataSourceV2Exec.scala:452)
at org.apache.spark.sql.execution.datasources.v2.V2TableWriteExec.$anonfun$writeWithV2$2(WriteToDataSourceV2Exec.scala:360)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:131)
at org.apache.spark.executor.Executor$TaskRunner.$anonfun$run$3(Executor.scala:497)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1439)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:500)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
... 1 more
-------
--
This is an automated message from the Apache Git Service.
To respond to the message, please log on to GitHub and use the
URL above to go to the specific comment.
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For queries about this service, please contact Infrastructure at:
users@infra.apache.org
---------------------------------------------------------------------
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For additional commands, e-mail: issues-help@iceberg.apache.org
[GitHub] [iceberg] RussellSpitzer edited a comment on issue #1894: Spark writes the Iceberg dual partition table to report an error
Posted by GitBox <gi...@apache.org>.
RussellSpitzer edited a comment on issue #1894:
URL: https://github.com/apache/iceberg/issues/1894#issuecomment-774303754
All Iceberg tables exist in the context of a Catalog. If you haven't specified one and just provided a path it is using the implied "HadoopCatalog". The table will have It's own Metadata.json file which will specify the schema of the table and partition spec. Once created, that is the schema and partition spec until it is specifically altered.
So in your example "./data-iceberg" will be treated as a HadoopTable and look in that directory for a Metadata.json with a specified schema and partitioning. If such a file did not exist you would get an error like
```scala
spark.read.parquet("/Users/russellspitzer/Temp/parquet_data").write.format("iceberg").save("/Users/russellspitzer/Temp/iceberg_table")
org.apache.iceberg.exceptions.NoSuchTableException: Table does not exist at location: /Users/russellspitzer/Temp/iceberg_table
```
Since you didn't get such an error means that a table does exist there and it has a schema.
When attempting attempting to write Spark saw "partitionBy" "a", "b", and preformed a shuffle on the input data
So all spark partitions are now like
```
SparkPartition 1 - > (1,1,1,1) , (1,1,2,1), (1,1,1,3) ...
SparkPartition 2 -> (1,2,1,1), (1,2,2,1), (1,2,1,2) ...
SparkPartition 3 -> (2,4,1,1), (2,4,2,1), (2,4,1,2) ...
```
The first two values are always the same within the spark partition, but the last two are in a random order.
Then the Iceberg writer gets one of these partitions and it creates a writer based on it's Metadata.json which let's say is not
the same as the `partitionedBy` argument. For this example let's say the table is actually partitioned by 'c'
By default in Spark Iceberg's writer opens a file for each partition assumes it will write all the values for that partition, then close the file. If it tries to write partition 1 it ends up doing the following
```
(1,1,1,1) -> Open partition c = 1 and start new file
(1,1,2,1) -> Close partition c = 1 file; Open partition c =2 and start new file
(1,1,1,3) -> Open partition c = 1 /// Error already closed file for Partition c=1
```
Since this is a runtime check you won't see this error with a single row since that will always be able to open and close the file without incident, but a dataset with multiple rows can potentially trigger this. Most of the time you want this Spark partitioning to match the Iceberg partitioning because this minimizes the number of files you need to actually write.
If you really don't want to have the Spark Partitioning match for some reason you can use the sortWithinPartitions to order the data within the partitions by their Iceberg partition values.
The other possibility is to use the new spark fanout writer which keeps all of the files open until they entire task has been completed.
Neither of these solutions is really as good as having your data partitioned like the data is expecting since that minimizes the number of files being written since all the files for a given partition will be done by the same Spark Task.
An example
```scala
scala> spark.sql("CREATE table ice.default.part (x Int, y Int) USING iceberg partitioned by (x) location '/Users/russellspitzer/Temp/iceberg_table'");
val df = spark.createDataFrame(Seq((1,1),(1,2),(1,3),(2,1),(2,2),(2,3),(3,1),(3,2),(3,3))).toDF("x","y")
scala> df.write.format("iceberg").partitionBy("x").mode("overwrite").saveAsTable("ice.default.part")
// No Error
scala> df.write.format("iceberg").partitionBy("y").mode("overwrite").saveAsTable("ice.default.part")
// Many errors
// Caused by: java.lang.IllegalStateException: Already closed files for partition: y=1
```
----------------------------------------------------------------
This is an automated message from the Apache Git Service.
To respond to the message, please log on to GitHub and use the
URL above to go to the specific comment.
For queries about this service, please contact Infrastructure at:
users@infra.apache.org
---------------------------------------------------------------------
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For additional commands, e-mail: issues-help@iceberg.apache.org
[GitHub] [iceberg] RussellSpitzer edited a comment on issue #1894: Spark writes the Iceberg dual partition table to report an error
Posted by GitBox <gi...@apache.org>.
RussellSpitzer edited a comment on issue #1894:
URL: https://github.com/apache/iceberg/issues/1894#issuecomment-774307653
And here is an example
```scala
scala> spark.sql("CREATE table ice.default.part (x Int, y Int) USING iceberg partitioned by (x) location '/Users/russellspitzer/Temp/iceberg_table'");
val df = spark.createDataFrame(Seq((1,1),(1,2),(1,3),(2,1),(2,2),(2,3),(3,1),(3,2),(3,3))).toDF("x","y")
scala> df.write.format("iceberg").partitionBy("x").mode("overwrite").saveAsTable("ice.default.part")
// No Error
scala> df.write.format("iceberg").partitionBy("y").mode("overwrite").saveAsTable("ice.default.part")
// Many errors
// Caused by: java.lang.IllegalStateException: Already closed files for partition: y=1
```
----------------------------------------------------------------
This is an automated message from the Apache Git Service.
To respond to the message, please log on to GitHub and use the
URL above to go to the specific comment.
For queries about this service, please contact Infrastructure at:
users@infra.apache.org
---------------------------------------------------------------------
To unsubscribe, e-mail: issues-unsubscribe@iceberg.apache.org
For additional commands, e-mail: issues-help@iceberg.apache.org